5. ADI forward modeling of point sources¶
Authors: Valentin Christiaens and Carlos Alberto Gomez GonzalezSuitable for VIP v1.0.0 onwardsLast update: 2022/09/21
Table of contents
- 5.1. Loading ADI data
- 5.2. Generating and injecting synthetic planets
- 5.3. Flux and position estimation with NEGFC
This tutorial shows:
- how to generate and inject fake companions in a cube;
- how to estimate the astrometry and photometry of a directly imaged companion, and associated uncertainties.
Let’s first import a couple of external packages needed in this tutorial:
[1]:
%matplotlib inline
from hciplot import plot_frames, plot_cubes
from matplotlib.pyplot import *
from matplotlib import pyplot as plt
import numpy as np
from packaging import version
In the following box we import all the VIP routines that will be used in this tutorial. The path to some routines has changed between versions 1.0.3 and 1.1.0, which saw a major revamp of the modular architecture, hence the if statements.
[2]:
import vip_hci as vip
vvip = vip.__version__
print("VIP version: ", vvip)
if version.parse(vvip) < version.parse("1.0.0"):
msg = "Please upgrade your version of VIP"
msg+= "It should be 1.0.0 or above to run this notebook."
raise ValueError(msg)
elif version.parse(vvip) <= version.parse("1.0.3"):
from vip_hci.conf import VLT_NACO
from vip_hci.metrics import cube_inject_companions
from vip_hci.negfc import (confidence, firstguess, mcmc_negfc_sampling, show_corner_plot, show_walk_plot,
speckle_noise_uncertainty)
from vip_hci.pca import pca, pca_annular, pca_annulus, pca_grid
else:
from vip_hci.config import VLT_NACO
from vip_hci.fm import (confidence, cube_inject_companions, cube_planet_free, firstguess, mcmc_negfc_sampling,
normalize_psf, show_corner_plot, show_walk_plot, speckle_noise_uncertainty)
from vip_hci.psfsub import median_sub, pca, pca_annular, pca_annulus, pca_grid
# common to all versions:
from vip_hci.fits import open_fits, write_fits, info_fits
from vip_hci.metrics import contrast_curve, detection, significance, snr, snrmap, throughput
from vip_hci.preproc import frame_crop
from vip_hci.var import fit_2dgaussian, frame_center
VIP version: 1.2.4
5.1. Loading ADI data¶
In the ‘dataset’ folder of the VIP_extras repository you can find a toy ADI (Angular Differential Imaging) cube and a NACO point spread function (PSF) to demonstrate the capabilities of VIP. This is an L’-band VLT/NACO dataset of beta Pictoris published in Absil et al. (2013) obtained using the Annular Groove Phase Mask (AGPM) Vortex coronagraph. The sequence has been heavily sub-sampled temporarily to make it
smaller. The frames were also cropped to the central 101x101 area. In case you want to plug-in your cube just change the path of the following cells.
More info on this dataset, and on opening and visualizing fits files with VIP in general, is available in Tutorial 1. Quick start.
Let’s load the data:
[3]:
psfnaco = '../datasets/naco_betapic_psf.fits'
cubename = '../datasets/naco_betapic_cube_cen.fits'
angname = '../datasets/naco_betapic_pa.fits'
cube = open_fits(cubename)
psf = open_fits(psfnaco)
pa = open_fits(angname)
Fits HDU-0 data successfully loaded. Data shape: (61, 101, 101)
Fits HDU-0 data successfully loaded. Data shape: (39, 39)
Fits HDU-0 data successfully loaded. Data shape: (61,)
[4]:
derot_off = 104.84 # NACO derotator offset for this observation (Absil et al. 2013)
TN = -0.45 # Position angle of true north for NACO at the epoch of observation (Absil et al. 2013)
angs = pa+derot_off+TN
Let’s measure the FWHM by fitting a 2D Gaussian to the core of the unsaturated non-coronagraphic PSF:
[5]:
%matplotlib inline
DF_fit = fit_2dgaussian(psf, crop=True, cropsize=9, debug=True, full_output=True)
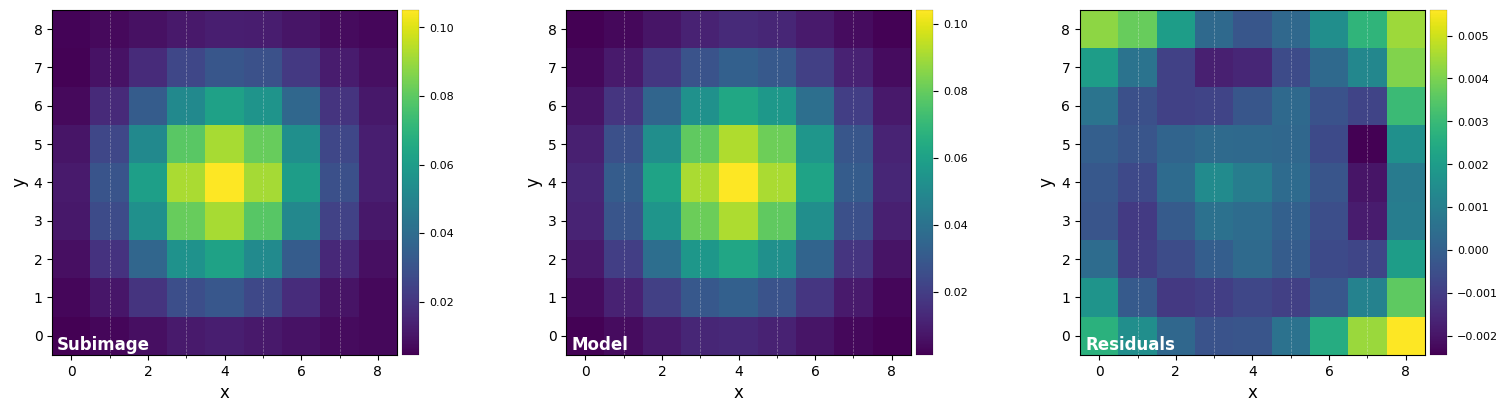
FWHM_y = 4.733218722257407
FWHM_x = 4.473682405059958
centroid y = 19.006680059041216
centroid x = 18.999424475165455
centroid y subim = 4.006680059041214
centroid x subim = 3.9994244751654535
amplitude = 0.10413004853269707
theta = -34.08563676836685
[6]:
fwhm_naco = np.mean([DF_fit['fwhm_x'],DF_fit['fwhm_y']])
print(fwhm_naco)
4.603450563658683
Let’s normalize the flux to one in a 1xFWHM aperture and crop the PSF array:
[7]:
psfn = normalize_psf(psf, fwhm_naco, size=19, imlib='ndimage-fourier')
Flux in 1xFWHM aperture: 1.228
[8]:
plot_frames(psfn, grid=True, size_factor=4)
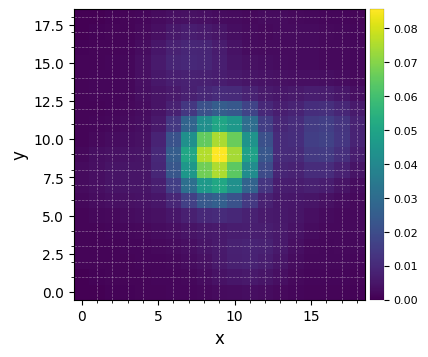
Let’s finally define the pixel scale for NACO (L’ band), which we get from a dictionary stored in the config subpackage:
[9]:
pxscale_naco = VLT_NACO['plsc']
print(pxscale_naco, "arcsec/px")
0.02719 arcsec/px
5.2. Generating and injecting synthetic planets¶
We first select an image library imlib for image operations (shifts, rotations) and associated interpolation order. ‘vip-fft’ is more accurate, but ‘skimage’ is faster, and ‘opencv’ even faster - see Tutorial 7 for more details.
[10]:
imlib_rot = 'opencv' #'skimage'
interpolation='lanczos4' #'biquintic'
The cube_inject_companions function in the fm module (VIP versions >= 1.1.0) makes the injection of fake companions at arbitrary fluxes and locations very easy. The normalized non-coronagraphic PSF should be provided for the injection. If the user does not have access to an observed PSF, the create_synth_psf from the var module can be used to create synthetic ones (based on 2D Gaussian, Moffat or Airy models).
Some procedures, e.g. the negative fake companion technique and the contrast curve generation, heavily rely on the injection of fake companions. The coordinates for the injection should be provided in the derotated image, while the actual injection occurs in the images of the input cube, i.e. in the rotated field.
[11]:
rad_fc = 30.5
theta_fc = 240
flux_fc = 400.
gt = [rad_fc, theta_fc, flux_fc]
[12]:
cubefc = cube_inject_companions(cube, psf_template=psfn, angle_list=angs, flevel=flux_fc, plsc=pxscale_naco,
rad_dists=[rad_fc], theta=theta_fc, n_branches=1,
imlib=imlib_rot, interpolation=interpolation)
Let’s set the corresponding cartesian coordinates:
[13]:
cy, cx = frame_center(cube[0])
x_fc = cx + rad_fc*np.cos(np.deg2rad(theta_fc))
y_fc = cy + rad_fc*np.sin(np.deg2rad(theta_fc))
xy_test = (x_fc, y_fc)
print('({:.1f}, {:.1f})'.format(xy_test[0],xy_test[1]))
(34.7, 23.6)
Let’s double-check the fake companion was injected at the right location, by post-processing the cube and checking the final image. Let’s use PCA, and infer the optimal \(n_{\rm pc}\) while we are at it - this will be useful for the next section.
[14]:
res_ann_opt = pca_grid(cubefc, angs, fwhm=fwhm_naco, range_pcs=(1,41,1), source_xy=xy_test, mode='annular',
annulus_width=4*fwhm_naco, imlib=imlib_rot, interpolation=interpolation,
full_output=True, plot=True)
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Starting time: 2022-05-04 13:37:46
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Done SVD/PCA with numpy SVD (LAPACK)
Running time: 0:00:00.057389
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Number of steps 41
Optimal number of PCs = 10, for S/N=13.415
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Coords of chosen px (X,Y) = 34.7, 23.6
Flux in a centered 1xFWHM circular aperture = 104.785
Central pixel S/N = 18.264
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Inside a centered 1xFWHM circular aperture:
Mean S/N (shifting the aperture center) = 13.415
Max S/N (shifting the aperture center) = 18.472
stddev S/N (shifting the aperture center) = 4.088
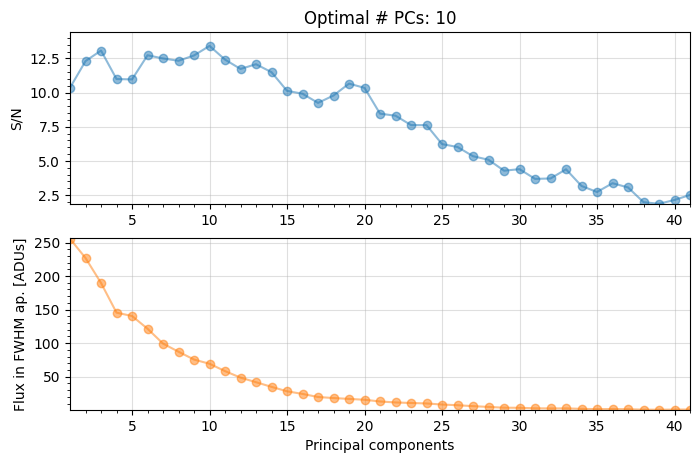
The grid search looking for the optimal number of principal components (npc) found that 10 principal components maximizes the S/N ratio of the injected fake companion.
[15]:
_, final_ann_opt, _, opt_npc_ann = res_ann_opt
[16]:
plot_frames(final_ann_opt, label='PCA-ADI annulus (opt. npc={:.0f})'.format(opt_npc_ann),
dpi=100, vmin=-5, colorbar=True, circle=xy_test)

We can see that the fake companion was indeed injected at the requested location.
5.3. Flux and position estimation with NEGFC¶
When a companion candidate is detected, the next step is to characterize it, i.e. infer its exact position (astrometry) and flux (photometry).
Question 5.1: Why would a simple 2D Gaussian fit (as performed e.g. for the stellar PSF in Section 5.1) be inappropriate to extract the astrometry and photometry of a candidate companion?
VIP implements the Negative fake companion (NEGFC) technique for robust extraction of the position and flux of detected point-like sources. The technique can be summarized as follow (see full description in Wertz et al. 2017):
- Estimate the position and flux of the planet, from either the visual inspection of reduced images or a previous estimator (see ABC below).
- Scale (in flux) and shift the normalized off-axis PSF to remove the estimate from the input data cube.
- Process the cube with PCA in a single annulus encompassing the point source.
- Measure residuals in an aperture centered on the approximate location of the companion candidate.
- Iterate on the position and flux of the injected negative PSF (steps 2-4), until the absolute residuals in the aperture are minimized (i.e. the injected negative companion flux and the position match exactly that of the true companion).
Iterations between steps 2-4 can be performed in one of 3 ways - sorted in increasing computation time and accuracy:
- a grid search on the flux only, provided a fixed estimate of the position (implemented in the
firstguessfunction); - a Nelder-Mead simplex algorithm (
firstguessfunction with thesimplex=Trueoption); - an MCMC sampler, which has the advantage to also yield uncertainties on each of the parameters of the point source (
mcmc_negfc_samplingfunction).
Different figures of merit can be used for minimization of the residuals (Wertz et al. 2017; Christiaens et al. 2021):
where \(j \in {1,...,N}\), \(N\) is the total number of pixels contained in the circular aperture around the companion candidate, \(\mu\) and \(\sigma\) are the mean and standard deviation (\(N_{\rm resel}\) degrees of freedom) of pixel intensities in a truncated annulus at the radius of the companion candidate, but avoiding the azimuthal region encompassing the negative side lobes.
5.3.1. Nelder-Mead based optimization¶
With the function firstguess, we can obtain a first estimation of the flux and position by running A) a naive grid minimization (grid of values for the flux through parameter f_range), and B) a Nelder-mead based minimization (if the parameter simplex is set to True). The latter is done based on the preliminary guess of the grid minimization. The maximum number of iterations and error can be set with the parameter simplex_options as a dicitionary (see scipy.minimize function
for the Nelder-Mead options).
Fisrt we define the position of the sources by examining a flux frame or S/N map. planets_xy_coord takes a list or array of X,Y pairs like ((x1,y1),(x2,y2)…(x_n,y_n)). Let’s take the coordinates of the previously injected companion.
Let’s test the algorithm with different values for the # of PCs: 5 and 25.
[17]:
r_lo, theta_lo, f_lo = firstguess(cubefc, angs, psfn, ncomp=5, planets_xy_coord=[xy_test],
fwhm=fwhm_naco, f_range=None, annulus_width=4*fwhm_naco,
aperture_radius=2, simplex=True, imlib=imlib_rot,
interpolation=interpolation, plot=True, verbose=True)
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Starting time: 2022-05-04 13:37:50
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Planet 0
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Planet 0: flux estimation at the position [34.749999999999986,23.58622518457463], running ...
Step | flux | chi2r
1/30 0.100 1.975
2/30 0.149 1.974
3/30 0.221 1.974
4/30 0.329 1.972
5/30 0.489 1.971
6/30 0.728 1.969
7/30 1.083 1.965
8/30 1.610 1.960
9/30 2.395 1.953
10/30 3.562 1.942
11/30 5.298 1.926
12/30 7.880 1.902
13/30 11.721 1.866
14/30 17.433 1.813
15/30 25.929 1.733
16/30 38.566 1.615
17/30 57.362 1.446
18/30 85.317 1.216
19/30 126.896 0.926
20/30 188.739 0.554
21/30 280.722 0.179
22/30 417.532 0.047
23/30 621.017 0.777
24/30 923.671 3.817
25/30 1373.824 11.783

Planet 0: preliminary position guess: (r, theta)=(30.5, 240.0)
Planet 0: preliminary flux guess: 417.5
Planet 0: Simplex Nelder-Mead minimization, running ...
Planet 0: Success: True, nit: 97, nfev: 224, chi2r: 0.030578838959905028
message: Optimization terminated successfully.
Planet 0: simplex result: (r, theta, f)=(30.535, 239.976, 383.055) at
(X,Y)=(34.72, 23.56)
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
DONE !
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Running time: 0:00:23.468048
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
[18]:
r_hi, theta_hi, f_hi = firstguess(cubefc, angs, psfn, ncomp=25, planets_xy_coord=[xy_test],
fwhm=fwhm_naco, f_range=None, annulus_width=4*fwhm_naco,
aperture_radius=2, imlib=imlib_rot, interpolation=interpolation,
simplex=True, plot=True, verbose=True)
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Starting time: 2022-05-04 13:38:14
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Planet 0
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Planet 0: flux estimation at the position [34.749999999999986,23.58622518457463], running ...
Step | flux | chi2r
1/30 0.100 0.295
2/30 0.149 0.295
3/30 0.221 0.295
4/30 0.329 0.295
5/30 0.489 0.295
6/30 0.728 0.295
7/30 1.083 0.295
8/30 1.610 0.295
9/30 2.395 0.295
10/30 3.562 0.294
11/30 5.298 0.293
12/30 7.880 0.292
13/30 11.721 0.291
14/30 17.433 0.290
15/30 25.929 0.286
16/30 38.566 0.283
17/30 57.362 0.275
18/30 85.317 0.249
19/30 126.896 0.223
20/30 188.739 0.180
21/30 280.722 0.107
22/30 417.532 0.058
23/30 621.017 0.191
24/30 923.671 0.422
25/30 1373.824 0.510
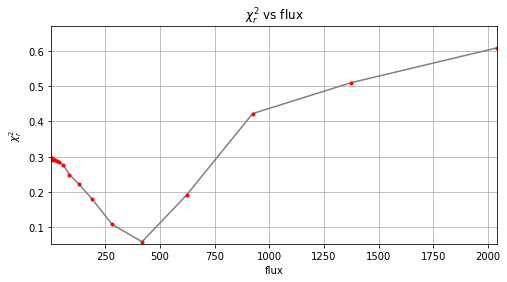
Planet 0: preliminary position guess: (r, theta)=(30.5, 240.0)
Planet 0: preliminary flux guess: 417.5
Planet 0: Simplex Nelder-Mead minimization, running ...
Planet 0: Success: True, nit: 71, nfev: 186, chi2r: 0.05277664606807796
message: Optimization terminated successfully.
Planet 0: simplex result: (r, theta, f)=(30.186, 239.897, 424.156) at
(X,Y)=(34.86, 23.89)
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
DONE !
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Running time: 0:00:19.281336
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
For both the \(n_{\rm pc} = 5\) and \(n_{\rm pc} = 25\) cases, the parameters estimated by the Nelder-Mead optimization are not exactly equal to the original values (radius=30.5, theta=240, flux=400), which reflects:
- the limitations of this heuristic minization procedure (depending on the initial guess the minimization may get trapped in a different local minimum);
- the higher residual speckle noise level in images obtained with low \(n_{\rm pc}\) values;
- the higher self-subtraction for high \(n_{\rm pc}\) values.
These estimates are provided without uncertainties (error bars). We will come back to this question later on.
For comparison, let’s use the optimal \(n_{\rm pc} = 10\) found in Section 5.2:
[19]:
r_0, theta_0, f_0 = firstguess(cubefc, angs, psfn, ncomp=opt_npc_ann, planets_xy_coord=[xy_test],
fwhm=fwhm_naco, f_range=None, annulus_width=4*fwhm_naco,
aperture_radius=2, imlib=imlib_rot, interpolation=interpolation,
simplex=True, plot=True, verbose=True)
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Starting time: 2022-05-04 13:38:33
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Planet 0
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Planet 0: flux estimation at the position [34.749999999999986,23.58622518457463], running ...
Step | flux | chi2r
1/30 0.100 1.992
2/30 0.149 1.992
3/30 0.221 1.991
4/30 0.329 1.990
5/30 0.489 1.989
6/30 0.728 1.987
7/30 1.083 1.984
8/30 1.610 1.980
9/30 2.395 1.974
10/30 3.562 1.964
11/30 5.298 1.951
12/30 7.880 1.931
13/30 11.721 1.900
14/30 17.433 1.855
15/30 25.929 1.785
16/30 38.566 1.696
17/30 57.362 1.554
18/30 85.317 1.315
19/30 126.896 1.031
20/30 188.739 0.650
21/30 280.722 0.224
22/30 417.532 0.056
23/30 621.017 0.656
24/30 923.671 2.701
25/30 1373.824 6.644
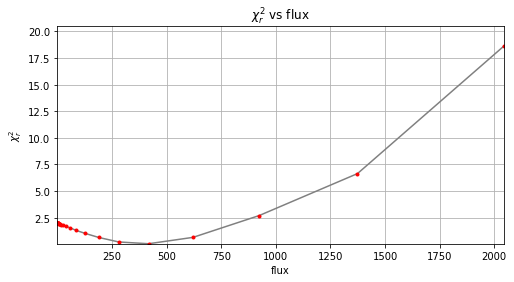
Planet 0: preliminary position guess: (r, theta)=(30.5, 240.0)
Planet 0: preliminary flux guess: 417.5
Planet 0: Simplex Nelder-Mead minimization, running ...
Planet 0: Success: True, nit: 79, nfev: 197, chi2r: 0.047533046198255234
message: Optimization terminated successfully.
Planet 0: simplex result: (r, theta, f)=(30.537, 239.907, 392.680) at
(X,Y)=(34.69, 23.58)
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
DONE !
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Running time: 0:00:20.609431
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
We see that using the optimal \(n_{\rm pc}\) leads to a closer parameter estimates to the ground truth.
Answer 5.1: If relying on a 2D Gaussian fit, both the photometry and astrometry would be biased by self-subtraction and the negative side lobes.
5.3.2. Planet subtraction¶
Let’s use the values obtained with the simplex optimization to subtract the planet with the function cube_planet_free.
First we define a list with the parameters (r, theta, flux) is each companion that we obtained via the NegFC, in this case one:
[20]:
plpar_fc = [(r_0[0], theta_0[0], f_0[0])]
Note: r_0, theta_0 and f_0 have the same length as the number of planet coordinates planets_xy_coord provided to firstguess. Here there is only one planet, so we take the zeroth index. The number of tuples in (i.e the length of) plpar_fc should match the number of planets.
[21]:
cube_emp = cube_planet_free(plpar_fc, cubefc, angs, psfn, imlib=imlib_rot, interpolation=interpolation)
Let’s double-check the fake companion was well removed by computing a PCA post-processed image:
[22]:
from vip_hci.preproc import cube_derotate
from vip_hci.config import time_ini, timing
t0 = time_ini()
for i in range(100):
fr_pca_emp = pca_annulus(cube_emp, angs, ncomp=opt_npc_ann, annulus_width=4*fwhm_naco,
r_guess=rad_fc, imlib=imlib_rot, interpolation=interpolation)
#cube_tmp = cube_derotate(cube_emp, angs, imlib=imlib_rot, interpolation=interpolation, edge_blend='')
timing(t0)
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Starting time: 2022-05-04 13:38:54
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Running time: 0:00:09.778283
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Let’s take a look at the PSF of the planet in the full-frame PCA final image and the same PSF in the frame resulting of processing the planet-subtracted cube:
[23]:
cropped_frame1 = frame_crop(final_ann_opt, cenxy=xy_test, size=15)
New shape: (15, 15)
[24]:
cropped_frame2 = frame_crop(fr_pca_emp, cenxy=xy_test, size=15)
New shape: (15, 15)
Let’s use both 'mode=surface' and the default image mode of plot_frames to show the residuals in the vicinity of the companion:
[25]:
plot_frames((cropped_frame1, cropped_frame2), mode='surface', vmax=8)

[26]:
plot_frames((final_ann_opt, fr_pca_emp), vmin = float(np.amin(final_ann_opt)),
vmax= float(np.amax(final_ann_opt)), axis=False)
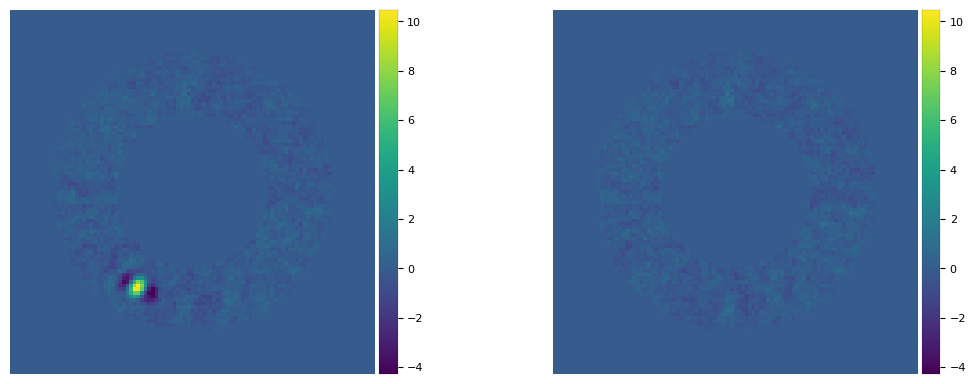
Not only the bright point-like signal is subtracted, but so are the negative side lobes. A subtraction not leaving any significant artifact/defect is a good sign that the inferred parameters are correct. However, keep in mind that even for slightly inaccurate parameters, the final image can still look relatively clean. Let’s take for example the parameters inferred with non-optimal \(n_{\rm pc}\):
[27]:
# planet parameters inferred from npc=5 used to make empty cube, then processed with PCA (npc=10)
plpar_fc = [(r_lo[0], theta_lo[0], f_lo[0])]
print(plpar_fc)
cube_emp = cube_planet_free(plpar_fc, cubefc, angs, psfn, imlib=imlib_rot, interpolation=interpolation)
final_ann_5 = pca_annulus(cubefc, angs, ncomp=5, annulus_width=4*fwhm_naco, r_guess=30.5,
imlib=imlib_rot, interpolation=interpolation)
fr_pca_emp5 = pca_annulus(cube_emp, angs, ncomp=5, annulus_width=4*fwhm_naco, r_guess=30.5,
imlib=imlib_rot, interpolation=interpolation)
plot_frames((final_ann_5, fr_pca_emp5), vmin = float(np.amin(final_ann_5)),
vmax= float(np.amax(final_ann_5)), axis=False)
[(30.534749178951024, 239.97617468137406, 383.05503327816507)]
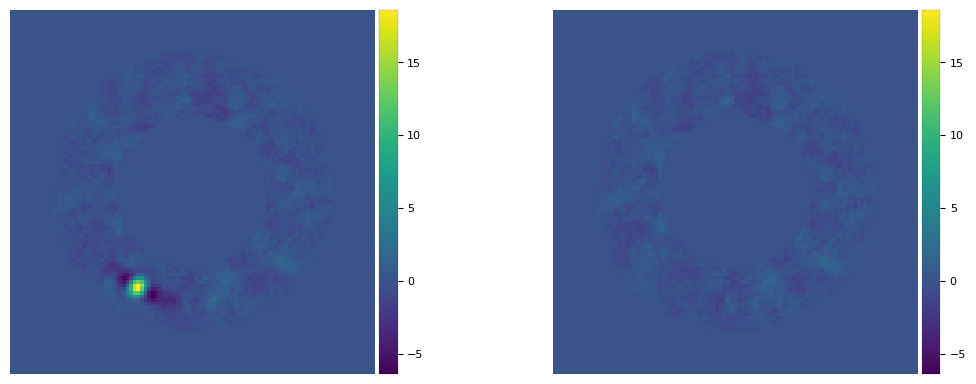
[28]:
# parameters inferred from npc=25 used to make empty cube, then processed with PCA (npc=10)
plpar_fc = [(r_hi[0], theta_hi[0], f_hi[0])]
print(plpar_fc)
cube_emp = cube_planet_free(plpar_fc, cubefc, angs, psfn, imlib=imlib_rot, interpolation=interpolation)
final_ann_25 = pca_annulus(cubefc, angs, ncomp=25, annulus_width=4*fwhm_naco, r_guess=30.5,
imlib=imlib_rot, interpolation=interpolation)
fr_pca_emp25 = pca_annulus(cube_emp, angs, ncomp=25, annulus_width=4*fwhm_naco, r_guess=30.5,
imlib=imlib_rot, interpolation=interpolation)
plot_frames((final_ann_25, fr_pca_emp25), vmin = float(np.amin(final_ann_25)),
vmax= float(np.amax(final_ann_25)), axis=False)
[(30.18557073057306, 239.89702835629325, 424.15580033622734)]

Inaccurate parameters still leading to an apparently good removal of the companion brings the question of the uncertainties on each of the three parameters characterizing the companion. The next sections are dedicated to this question.
5.3.3. NEGFC technique coupled with MCMC¶
5.3.3.1. Running the MCMC sampler¶
MCMC is a more robust way of obtaining the flux and position. It samples the posterior distributions of the parameters and from them we can infer both the most likely parameter values and uncertainties on each parameter. The relevant function is mcmc_negfc_sampling, which can accept a number of parameters. Let’s define them in the next few boxes:
Let’s first define observation-related parameters, such as the non-coronagraphic psf, its FWHM and the pixel scale od the detector:
[29]:
obs_params = {'psfn': psfn,
'fwhm': fwhm_naco}
In NEGFC, PCA in a single annulus is used by default to speed up the algorithm - although other algorithms can be used through the algo parameter. Let’s set the \(n_{\rm pc}\) to the optimal \(n_{\rm pc}\) found in Section 5.2. We set the width of the annulus on which PCA is performed (in pixels) with the annulus_width parameter. We also set a few other algorithm-related parameters in the following box:
[30]:
annulus_width = 4*fwhm_naco
algo_params = {'algo': pca_annulus,
'ncomp': opt_npc_ann,
'annulus_width': annulus_width,
'svd_mode': 'lapack',
'imlib': imlib_rot,
'interpolation': interpolation}
The choice of log-likelihood expression to be used is determined by mu_sigma and fm. If the former is True (default; mu and sigma calculated automatically) or a tuple of 2 values, the following figure of merit will be used: \(\chi^2 = \sum_j^N \frac{(I_j-\mu)^2}{\sigma^2}\). Otherwise, the choice will be determined by fm: ‘sum’ for the sum of absolute residuals, or ‘stddev’ for the standard deviation of residuals (which can be useful if the point source is contained within a more
extended signal). aperture_radius is the radius of the aperture (in fwhm units) in which the residual intensities \(I_j\) are considered.
[31]:
mu_sigma=True
aperture_radius=2
negfc_params = {'mu_sigma': mu_sigma,
'aperture_radius': aperture_radius}
Parameter initial_state corresponds to the initial first estimation of the planets parameters (r, theta, flux). We set it to the result of the simplex optimization, obtained with optimal \(n_{\rm pc}\). Note that the MCMC minimization can only run for one companion candidate at a time, hence the first dimension of init should always be 3 (not the number of planets, as opposed to planets_xy_coord in firstguess).
[32]:
initial_state = np.array([r_0[0], theta_0[0], f_0[0]])
Beware that the MCMC procedure is a CPU intensive procedure and can take several hours when run properly on a standard laptop. We use the affine invariant sampler from emcee which can be run in parallel (nproc sets the number of CPUs to be used). At least 100 walkers are recommended for our MCMC chain, although both the number of walkers and iterations will depend on your dataset. For the sake of preventing this tutorial to take too long to run, we set the maximum number of iterations to
500, although feel free to significantly increase it in case of non-convergence.
[33]:
from multiprocessing import cpu_count
nwalkers, itermin, itermax = (100, 200, 500)
mcmc_params = {'nwalkers': nwalkers,
'niteration_min': itermin,
'niteration_limit': itermax,
'bounds': None,
'nproc': cpu_count()//2}
Another update from Christiaens et al. (2021) is that the convergence can now be evaluated based on the auto-correlation time (see more details in the documentation of ``emcee` <https://emcee.readthedocs.io/en/stable/tutorials/autocorr/>`__), instead of the Gelman-Rubin test, which is inappropriate for non-independent samples (such as an MCMC chain). We set the convergence test to the autocorrelation time based criterion using conv_test='ac' (instead of Gelman-Rubin ‘gb’). We also set the
autocorrelation criterion \(N/\tau >= a_c\), where \(N\) is the number of samples and \(\tau\) the autocorrelation time, to ac_c=50 (the value recommended in the emcee documentation). Finally, we set the number of consecutive times the criterion must be met to: ac_count_thr=1, and the maximum gap in number of steps between 2 checks of the convergence criterion to: check_maxgap=50. If the maximum number of iterations niteration_limit is large enough, the chain will
stop upon meeting the convergence criterion (spoiler: it won’t be the case here since we chose a small value for niteration_limit).
[34]:
conv_test, ac_c, ac_count_thr, check_maxgap = ('ac', 50, 1, 50)
conv_params = {'conv_test': conv_test,
'ac_c': ac_c,
'ac_count_thr': ac_count_thr,
'check_maxgap': check_maxgap}
Setting bounds=None does not mean no bounds are used for parameter exploration, but rather that they are set automatically to:
- \(r \in [r_0-w_{ann}/2, r_0+w_{ann}/2]\), where \(w_{ann}\) is the
annulus_width, - \(\theta \in [\theta_0-\Delta {\rm rot}, \theta_0+\Delta {\rm rot}]\), where \(\Delta {\rm rot}\) is the angle subtended by min(
aperture_radius/2,``fwhm``) at \(r_0\), - \(f \in [0.1*f_0, 2*f_0]\),
where (\(r_0, \theta_0, f_0\)) = initial_state. If the bounds are provided manually (as a tuple of tuples), they will supersede the automatic setting above.
Now let’s start the sampler. Note that this step is computer intensive and may take a long time to run depending on your machine. Feel free to skip the next box if you do not wish to run the MCMC or can’t run it in a reasonable time. The results are already saved as ‘MCMC_results’ in the ‘datasets’ folder and will be loaded in the subsequent boxes.
[35]:
chain = mcmc_negfc_sampling(cubefc, angs, **obs_params, **algo_params, **negfc_params,
initial_state=initial_state, **mcmc_params, **conv_params,
display=True, verbosity=2, save=False, output_dir='./')
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
Starting time: 2022-05-04 13:39:04
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
MCMC sampler for the NEGFC technique
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
The mean and stddev in the annulus at the radius of the companion (excluding the PA area directly adjacent to it) are -0.01 and 1.48 respectively.
Beginning emcee Ensemble sampler...
emcee Ensemble sampler successful
Start of the MCMC run ...
Step | Duration/step (sec) | Remaining Estimated Time (sec)
0 3.47851 1735.77874
1 1.38965 692.04520
2 1.38548 688.58207
3 1.36664 677.85542
4 1.37508 680.66460
5 1.38777 685.55739
6 1.36152 671.22936
7 1.37935 678.63823
8 1.39976 687.28216
9 1.38258 677.46469
10 1.36537 667.66397
11 1.37168 669.37935
12 1.36878 666.59781
13 1.36647 664.10636
14 1.36966 664.28704
15 1.36997 663.06451
16 1.36980 661.61340
17 1.38879 669.39823
18 1.37246 660.15182
19 1.38434 664.48368
20 1.37649 659.34063
21 1.36439 652.18033
22 1.36600 651.58152
23 1.36397 649.24734
24 1.36953 650.52723
25 1.36554 647.26454
26 1.37040 648.19873
27 1.36910 646.21567
28 1.36674 643.73219
29 1.37244 645.04821
30 1.36953 642.31191
31 1.37786 644.83801
32 1.36838 639.03486
33 1.37587 641.15728
34 1.38366 643.40190
35 1.37418 637.61859
36 1.36562 632.28345
37 1.37518 635.33316
38 1.37097 632.01717
39 1.37377 631.93512
40 1.36616 627.06744
41 1.41404 647.63032
42 1.42133 649.54918
43 1.37764 628.20202
44 1.37153 624.04660
45 1.38171 627.29725
46 1.37053 620.85100
47 1.37285 620.52639
48 1.36444 615.36064
49 1.36433 613.94895
50 1.36821 614.32449
ac convergence test in progress...
51 1.75774 787.46618
52 1.37224 613.39083
53 1.36716 609.75158
54 1.36701 608.31945
55 1.37034 608.42963
56 1.36799 606.01824
57 1.36227 602.12511
58 1.37602 606.82438
59 1.37225 603.79000
60 1.36685 600.04496
61 1.36519 597.95497
62 1.35685 592.94476
63 1.36890 596.84084
64 1.36670 594.51493
65 1.37584 597.11499
66 1.36403 590.62542
67 1.36657 590.35824
68 1.36726 589.28906
69 1.36786 588.18109
70 1.35679 582.06377
71 1.36564 584.49349
72 1.36407 582.45618
73 1.36285 580.57410
74 1.35704 576.74115
75 1.36997 580.86940
76 1.38369 585.30214
77 1.38872 586.03984
78 1.39496 587.27816
79 1.39433 585.61944
80 1.38035 578.36749
81 1.39219 581.93542
82 1.37591 573.75447
83 1.37403 571.59440
84 1.37887 572.22897
85 1.46772 607.63442
86 1.38292 571.14761
87 1.36528 562.49618
88 1.36131 559.49800
89 1.37875 565.28586
90 1.37170 561.02407
91 1.36405 556.53118
92 1.37014 557.64495
93 1.36063 552.41416
94 1.36496 552.80799
95 1.35604 547.83895
96 1.37475 554.02224
97 1.35842 546.08323
98 1.36179 546.07699
99 1.37333 549.33040
100 1.38278 551.73002
ac convergence test in progress...

101 1.93721 771.00878
102 1.36099 540.31224
103 1.37314 543.76502
104 1.36566 539.43491
105 1.37684 542.47299
106 1.39130 546.78090
107 1.37662 539.63347
108 1.37140 536.21896
109 1.36952 534.11124
110 1.36918 532.61063
111 1.36899 531.16967
112 1.35915 525.98911
113 1.35839 524.33700
114 1.36695 526.27498
115 1.36556 524.37466
116 1.36960 524.55527
117 1.37089 523.68189
118 1.35611 516.67677
119 1.37855 523.85014
120 1.37659 521.72723
121 1.35964 513.94316
122 1.36840 515.88793
123 1.37461 516.85336
124 1.36725 512.71837
125 1.36842 511.78833
126 1.35909 506.94020
127 1.36349 507.21642
128 1.40657 521.83858
129 1.39825 517.35398
130 1.37142 506.05361
131 1.36641 502.83778
132 1.35998 499.11303
133 1.37224 502.23984
134 1.36263 497.36031
135 1.36428 496.59646
136 1.38187 501.61881
137 1.35364 490.01804
138 1.35730 489.98602
139 1.37474 494.90640
140 1.36030 488.34770
141 1.36739 489.52383
142 1.36742 488.16894
143 1.37003 487.73175
144 1.37192 487.03195
145 1.35985 481.38584
146 1.35557 478.51797
147 1.36797 481.52685
148 1.36882 480.45547
149 1.36526 477.84205
150 1.38094 481.94876
ac convergence test in progress...

151 1.75770 611.67786
152 1.36334 473.07933
153 1.36924 473.75739
154 1.36606 471.29104
155 1.36057 468.03608
156 1.37101 470.25540
157 1.36387 466.44491
158 1.36719 466.21077
159 1.36894 465.43858
160 1.36576 462.99366
161 1.36094 459.99907
162 1.36516 460.05926
163 1.37645 462.48821
164 1.37219 459.68466
165 1.35647 453.06031
166 1.36374 454.12475
167 1.35243 449.00610
168 1.37150 453.96617
169 1.37865 454.95516
170 1.37030 450.82771
171 1.39785 458.49513
172 1.43137 468.05701
173 1.37182 447.21299
174 1.36833 444.70757
175 1.35695 439.65245
176 1.37037 442.62854
177 1.36034 438.02819
178 1.36000 436.56000
179 1.36537 436.91840
180 1.37311 438.02337
181 1.36582 434.33012
182 1.37269 435.14368
183 1.36729 432.06490
184 1.36564 430.17597
185 1.37111 430.52854
186 1.34831 422.02259
187 1.35744 423.52128
188 1.39528 433.93332
189 1.36377 422.76808
190 1.36072 420.46186
191 1.35898 418.56522
192 1.36317 418.49411
193 1.38411 423.53888
194 1.36445 416.15786
195 1.35518 411.97320
196 1.37015 415.15515
197 1.36743 412.96537
198 1.35415 407.60005
199 1.35116 405.34650
200 1.35808 406.06562
ac convergence test in progress...

Auto-corr tau/N = [0.06893561 0.06749297 0.0631891 ]
tau/N <= 0.02 = [False False False]
201 1.77288 528.31913
202 1.37000 406.88881
203 1.37306 406.42458
204 1.36606 402.98711
205 1.36417 401.06569
206 1.36871 401.03203
207 1.37946 402.80320
208 1.36279 396.57131
209 1.36843 396.84499
210 1.37597 397.65446
211 1.36622 393.47050
212 1.36528 391.83536
213 1.36224 389.60121
214 1.36528 389.10395
215 1.43257 406.84931
216 1.43605 406.40187
217 1.36902 386.06279
218 1.36523 383.62935
219 1.35951 380.66364
220 1.37400 383.34656
221 1.35613 377.00386
222 1.36476 378.03907
223 1.35997 375.35282
224 1.37137 377.12757
225 1.35623 371.60757
226 1.36196 371.81617
227 1.36167 370.37560
228 1.36531 369.99955
229 1.37026 369.97047
230 1.36312 366.67794
231 1.35774 363.87432
232 1.36135 363.48045
233 1.36352 362.69499
234 1.36618 362.03664
235 1.36623 360.68393
236 1.36985 360.27055
237 1.37298 359.72024
238 1.36663 356.69147
239 1.36002 353.60442
240 1.36172 352.68626
241 1.36940 353.30546
242 1.37169 352.52356
243 1.37288 351.45856
244 1.36466 347.98932
245 1.36358 346.34907
246 1.36706 345.86643
247 1.36993 345.22160
248 1.37215 344.40890
249 1.36566 341.41525
250 1.36199 339.13501
ac convergence test in progress...
Auto-corr tau/N = [0.06364067 0.06439249 0.06246105]
tau/N <= 0.02 = [False False False]
251 1.94756 482.99538
252 1.36337 336.75140
253 1.36100 334.80649
254 1.37141 335.99520
255 1.37027 334.34564
256 1.37115 333.18945
257 1.36799 331.05406
258 1.44828 349.03452
259 1.44380 346.51224
260 1.36711 326.74001
261 1.37324 326.83041
262 1.36312 323.06062
263 1.36307 321.68428
264 1.37064 322.10087
265 1.35906 318.01910
266 1.36530 318.11467
267 1.36447 316.55727
268 1.36189 314.59728
269 1.38437 318.40625
270 1.36441 312.45081
271 1.35902 309.85565
272 1.37468 312.05191
273 1.36083 307.54781
274 1.36878 307.97482
275 1.36476 305.70714
276 1.35680 302.56662
277 1.36779 303.64938
278 1.37177 303.16051
279 1.36021 299.24664
280 1.36642 299.24598
281 1.36069 296.62977
282 1.36457 296.11256
283 1.36525 294.89378
284 1.36005 292.41139
285 1.36183 291.43141
286 1.36830 291.44747
287 1.36051 288.42770
288 1.36444 287.89768
289 1.36100 285.81084
290 1.36291 284.84882
291 1.36013 282.90787
292 1.36963 283.51403
293 1.36475 281.13891
294 1.35840 278.47262
295 1.36955 279.38881
296 1.37323 278.76630
297 1.36603 275.93766
298 1.36052 273.46392
299 1.35451 270.90260
300 1.37308 273.24232
ac convergence test in progress...
Auto-corr tau/N = [0.05879776 0.05591288 0.06160337]
tau/N <= 0.02 = [False False False]
301 1.80238 356.87124
302 1.44800 285.25659
303 1.42286 278.88036
304 1.36092 265.37999
305 1.35983 263.80624
306 1.36730 263.88967
307 1.36941 262.92595
308 1.36853 261.38866
309 1.36217 258.81325
310 1.36594 258.16190
311 1.37018 257.59403
312 1.36369 255.01003
313 1.36981 254.78429
314 1.35805 251.23907
315 1.35745 249.77154
316 1.38237 252.97298
317 1.38293 251.69253
318 1.37136 248.21598
319 1.36390 245.50290
320 1.36443 244.23351
321 1.37658 245.03053
322 1.36869 242.25902
323 1.37047 241.20202
324 1.35917 237.85405
325 1.36901 238.20739
326 1.37334 237.58799
327 1.37183 235.95562
328 1.35928 232.43603
329 1.37152 233.15823
330 1.36776 231.15212
331 1.37223 230.53414
332 1.35824 226.82541
333 1.37457 228.17912
334 1.36473 225.18095
335 1.37727 225.87244
336 1.36999 223.30756
337 1.36504 221.13599
338 1.37452 221.29740
339 1.37399 219.83856
340 1.36184 216.53256
341 1.37440 217.15473
342 1.35969 213.47180
343 1.36336 212.68369
344 1.36208 211.12256
345 1.36396 210.04969
346 1.48087 226.57342
347 1.36201 207.02537
348 1.32350 199.84850
349 1.34267 201.39990
350 1.32945 198.08850
ac convergence test in progress...
Auto-corr tau/N = [0.05431862 0.05251512 0.05584251]
tau/N <= 0.02 = [False False False]
351 1.75272 259.40256
352 1.34394 197.55977
353 1.34163 195.87856
354 1.34003 194.30435
355 1.34685 193.94611
356 1.32480 189.44626
357 1.32372 187.96796
358 1.32240 186.45812
359 1.32098 184.93706
360 1.34424 186.84964
361 1.32233 182.48195
362 1.34912 184.82930
363 1.32204 179.79717
364 1.32098 178.33216
365 1.33035 178.26690
366 1.36083 180.99052
367 1.38104 182.29702
368 1.35296 177.23815
369 1.35963 176.75164
370 1.32232 170.57876
371 1.32080 169.06291
372 1.31747 167.31907
373 1.34191 169.08079
374 1.32056 165.07025
375 1.34571 166.86779
376 1.35669 166.87336
377 1.35366 165.14664
378 1.32598 160.44370
379 1.32274 158.72928
380 1.32460 157.62764
381 1.36931 161.57905
382 1.32920 155.51640
383 1.32964 154.23789
384 1.35979 156.37608
385 1.36325 155.41004
386 1.32564 149.79777
387 1.32717 148.64326
388 1.35921 150.87242
389 1.35952 149.54687
390 1.42843 155.69843
391 1.45226 156.84397
392 1.32920 142.22472
393 1.32644 140.60243
394 1.34765 141.50357
395 1.33825 139.17758
396 1.32440 136.41351
397 1.33204 135.86839
398 1.33378 134.71218
399 1.33044 133.04370
400 1.32203 130.88137
ac convergence test in progress...

Auto-corr tau/N = [0.05007131 0.04780584 0.04992041]
tau/N <= 0.02 = [False False False]
401 1.94558 190.66704
402 1.35482 131.41773
403 1.34879 129.48432
404 1.33167 126.50894
405 1.33250 125.25491
406 1.32852 123.55255
407 1.32884 122.25346
408 1.33683 121.65144
409 1.32430 119.18727
410 1.34801 119.97316
411 1.32566 116.65764
412 1.32543 115.31224
413 1.32671 114.09740
414 1.35313 115.01605
415 1.37420 115.43297
416 1.36264 113.09895
417 1.35015 110.71214
418 1.32385 107.23169
419 1.32301 105.84048
420 1.35332 106.91267
421 1.31920 102.89776
422 1.34429 103.51025
423 1.35127 102.69629
424 1.32177 99.13260
425 1.32626 98.14294
426 1.33582 97.51493
427 1.32242 95.21410
428 1.32658 94.18725
429 1.33272 93.29040
430 1.31968 91.05820
431 1.32835 90.32746
432 1.37294 91.98664
433 1.32567 87.49415
434 1.39452 90.64354
435 1.42596 91.26157
436 1.33659 84.20511
437 1.35165 83.80242
438 1.34022 81.75366
439 1.34367 80.62038
440 1.33631 78.84241
441 1.35414 78.54024
442 1.33743 76.23368
443 1.34242 75.17546
444 1.34213 73.81715
445 1.35488 73.16330
446 1.34346 71.20317
447 1.33739 69.54444
448 1.33476 68.07291
449 1.35572 67.78575
450 1.33413 65.37247
ac convergence test in progress...
Auto-corr tau/N = [0.0461422 0.04651919 0.04701059]
tau/N <= 0.02 = [False False False]
451 1.78397 85.63051
452 1.34145 63.04829
453 1.33849 61.57077
454 1.34721 60.62454
455 1.34479 59.17063
456 1.37163 58.98030
457 1.35656 56.97544
458 1.34790 55.26402
459 1.33506 53.40248
460 1.33735 52.15646
461 1.31649 50.02654
462 1.33691 49.46552
463 1.32977 47.87183
464 1.36603 47.81091
465 1.34201 45.62824
466 1.34397 44.35098
467 1.34131 42.92189
468 1.36327 42.26140
469 1.33844 40.15329
470 1.34606 39.03560
471 1.34264 37.59400
472 1.35083 36.47246
473 1.34537 34.97962
474 1.36461 34.11512
475 1.35284 32.46811
476 1.33381 30.67763
477 1.33902 29.45851
478 1.40367 29.47703
479 1.47148 29.42952
480 1.38129 26.24453
481 1.35009 24.30162
482 1.34242 22.82122
483 1.34057 21.44912
484 1.34618 20.19270
485 1.35671 18.99397
486 1.35507 17.61594
487 1.33743 16.04915
488 1.35180 14.86977
489 1.34272 13.42715
490 1.37927 12.41345
491 1.34466 10.75728
492 1.35501 9.48506
493 1.33119 7.98712
494 1.34971 6.74856
495 1.34203 5.36812
496 1.35477 4.06430
497 1.33913 2.67826
498 1.33900 1.33900
499 1.34192 0.00000
We have reached the limit # of steps without convergence
Running time: 0:11:31.680492
――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――――
If you ran the previous box and wish to write your results, set write=True in the next box. This will pickle the MCMC chain.
[36]:
write=False
if write:
import pickle
output = {'chain':chain}
with open('../datasets/my_MCMC_results', 'wb') as fileSave:
pickle.dump(output, fileSave)
Note that an alternative to the box above, is to provide output_dir in the call to mcmc_negfc_sampling and set save to True. This will save the results as a pickle including additional keys: apart from ‘chain’, it will also save ‘input_parameters’, ‘AR’ (acceptance ratio), and ‘lnprobability’.
Pickled results can be loaded from disk like this:
[37]:
import pickle
with open('../datasets/MCMC_results','rb') as fi:
myPickler = pickle.Unpickler(fi)
mcmc_result = myPickler.load()
print(mcmc_result.keys())
chain = mcmc_result['chain']
dict_keys(['chain', 'input_parameters', 'AR', 'lnprobability'])
The most accurate approach would involve setting a large enough maximum number of iterations and using FFT-based rotation for PCA. The latter in particular, may change a bit the most likely parameter values given the better flux conservation. However, these changes would involve over ~3 orders of magnitude longer calculation time. It is therefore intractable for a personal laptop and not shown in this notebook. If you have access to a supercomputer feel free to adopt these changes though. The results after 500 iterations are nonetheless good enough for illustrative purpose:
5.3.3.2. Visualizing the MCMC chain: corner plots and walk plots¶
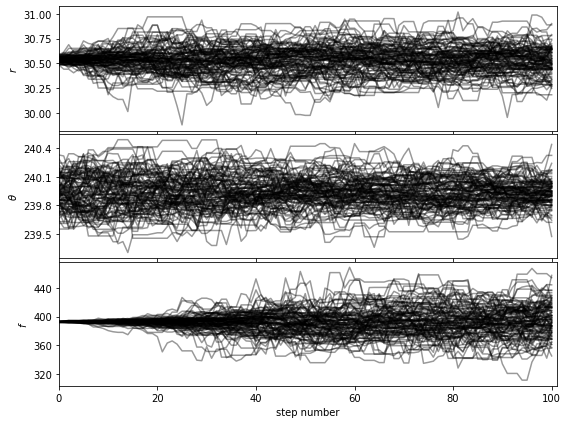
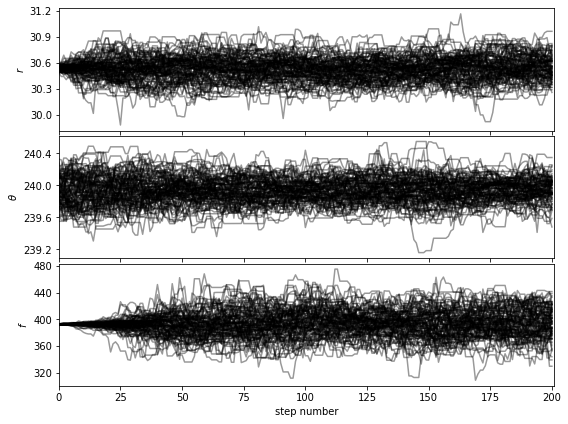
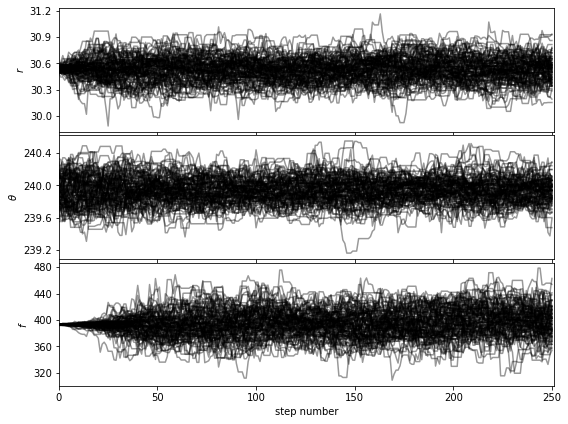
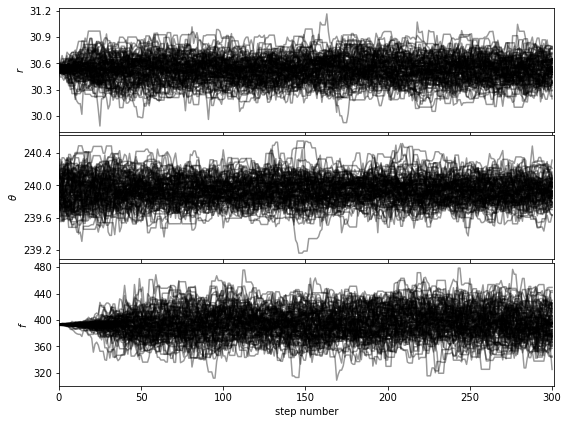
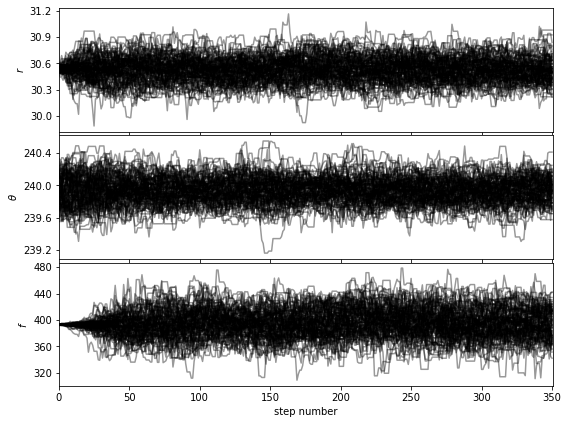
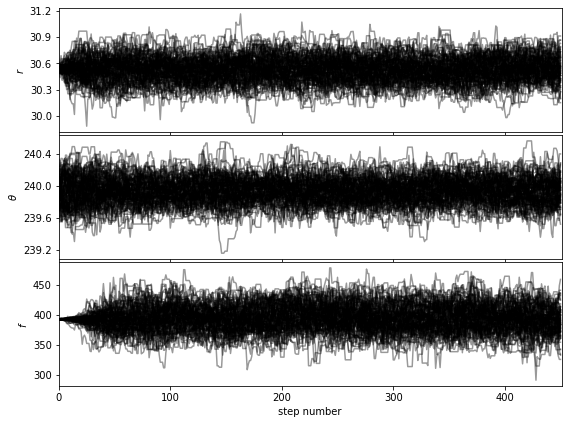
Let’s first check that the walk plots look ok:
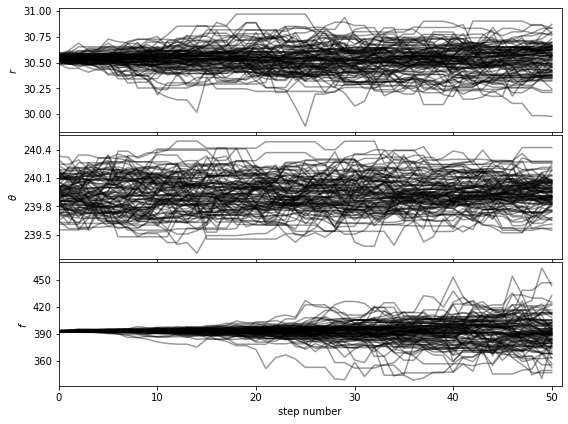
[38]:
show_walk_plot(chain)
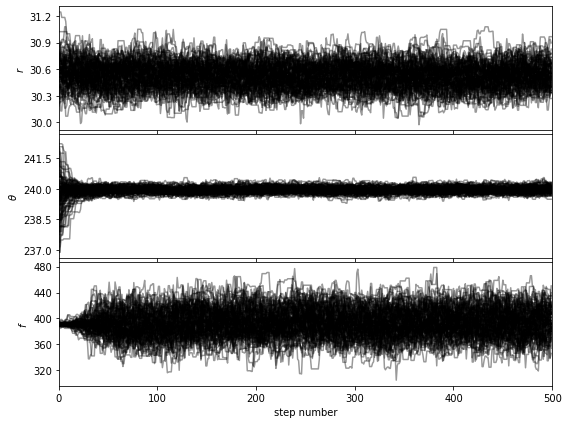
Then based on the walk plot, let’s burn-in the first 30% of the chain, to calculate the corner plots:
[39]:
burnin = 0.3
burned_chain = chain[:, int(chain.shape[1]//(1/burnin)):, :]
show_corner_plot(burned_chain)

For the purpose of this tutorial and to limit computation time we set the maximum number of iterations to 500 for 100 walkers. The look of the corner plots may improve with more samples (i.e. higher number of iterations, for a given burn-in ratio). This can be tested by setting the max. number of iterations arbitrarily high for the autocorrelation-time convergence criterion to be met.
5.3.3.3. Highly probable values and confidence intervals¶
Now let’s determine the most highly probable value for each model parameter, as well as the 1-sigma confidence interval. For this, let’s first flatten the chains:
[40]:
isamples_flat = chain[:, int(chain.shape[1]//(1/burnin)):, :].reshape((-1,3))
Then use the confidence function:
[41]:
val_max, conf = confidence(isamples_flat, cfd=68, gaussian_fit=False, verbose=True, save=False,
gt=gt, ndig=1, title=True)
percentage for r: 69.79202279202278%
percentage for theta: 69.02279202279202%
percentage for f: 68.94017094017094%
Confidence intervals:
r: 30.57489764243517 [-0.20421878422770945,0.08279139901123145]
theta: 239.91771004343246 [-0.14527599962792692,0.18317408648738365]
f: 390.7650582608513 [-20.054061636888093,25.285555976945886]
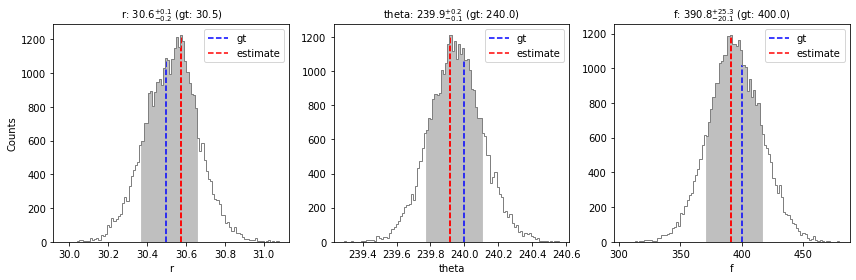
Using the confidence function with the gaussian_fit=True option, it is possible to fit a Gaussian to the posterior distribution of each parameter, and infer associated uncertainty values.
[42]:
mu, sigma = confidence(isamples_flat, cfd=68, gaussian_fit=True, verbose=True, save=False,
gt=gt, ndig=1, title=True)
percentage for r: 69.79202279202278%
percentage for theta: 69.02279202279202%
percentage for f: 68.94017094017094%
Confidence intervals:
r: 30.57489764243517 [-0.20421878422770945,0.08279139901123145]
theta: 239.91771004343246 [-0.14527599962792692,0.18317408648738365]
f: 390.7650582608513 [-20.054061636888093,25.285555976945886]
Gaussian fit results:
r: 30.525747462344565 +-0.13676103149725966
theta: 239.94793373875024 +-0.15744912409220196
f: 394.1338634773366 +-21.669634439677626

It is recommended to report the results as confidence intervals (i.e. with possibly asymmetric uncertainties) as long as the bin interval is small enough. Here, we also fitted the residual posterior distribution of each parameter to a Gaussian distribution (this shape is the expected one if the noise has been well whitened, but is not necessarily guaranteed at all separations depending on the adopted \(n_{\rm pc}\)). In case, these distributions look Gaussian, the inferred \(\sigma\) value may be a more accurate uncertainty value for the different parameters.
We can see that the confidence intervals inferred by NEGFC for the different parameters encompass the ground truth values used for injection (in particular for the flux).
Note that depending on your choice of mu_sigma, you may have to calculate separately another source of uncertainty. Indeed, with the original expression for \(\chi^2\) (Wertz et al. 2017; mu_sigma=False), only the uncertainty associated to photon noise was reflected in the MCMC results. With the new expression (mu_sigma=True), the \(\chi^2\) expression also takes into account the residual speckle noise at the radial separation of the companion candidate. Our tests suggest
that similar final uncertainties can be obtained in either of these 2 ways:
- the uncertainties obtained with MCMC when setting
mu_sigma=True; - the uncertainties obtained by combining quadratically the uncertainties obtained with MCMC (setting
mu_sigma=Falseandfm = 'sum'), and the residual speckle uncertainties inferred as in Sec. 5.3.4 (Wertz et al. 2017).
5.3.4. Residual speckle uncertainty¶
Only needed if using ``mu_sigma=False`` in your call to mcmc_negfc_sampling!
Residual speckle noise can also bias the best parameter estimates found for the companion (if not taken into account in the MCMC). To evaluate the uncertainty associated to this additional source of noise, it is recommended to inject a large number of fake companions at the same radius and flux as estimated for the true companion but different azimuths, and then estimate their parameters using simplex-NEGFC. The distribution of differences with respect to injected parameters can then give us an
idea of the residual speckle noise uncertainty. This is done in VIP with the speckle_noise_uncertainty function (see also Sec. 3.3 in Wertz et al. 2017 for more details).
Let’s use the planet parameters inferred by the MCMC-NEGFC algorithm:
[43]:
pl_par = (val_max['r'],val_max['theta'],val_max['f'])
pl_par
[43]:
(30.57489764243517, 239.91771004343246, 390.7650582608513)
[44]:
algo_options={'ncomp':opt_npc_ann, 'annulus_width':4*fwhm_naco, 'imlib':imlib_rot,
'interpolation':interpolation}
speckle_res = speckle_noise_uncertainty(cubefc, pl_par, np.linspace(0,359,360), angs, pca_annulus,
psfn, fwhm_naco, aperture_radius=2, fmerit='sum',
algo_options=algo_options, transmission=None, mu_sigma=None,
wedge=None, weights=None, force_rPA=False, nproc=None,
simplex_options=None, bins=None, save=False, output=None,
verbose=True, full_output=True, plot=True)
#######################################################
### SPECKLE NOISE DETERMINATION ###
#######################################################
Number of steps: 360
Process is running for angle: 0.00
Process is running for angle: 18.00
Process is running for angle: 36.00
Process is running for angle: 54.00
Process is running for angle: 72.00
Process is running for angle: 1.00
Process is running for angle: 37.00
Process is running for angle: 19.00
Process is running for angle: 73.00
Process is running for angle: 55.00
Process is running for angle: 38.00
Process is running for angle: 2.00
Process is running for angle: 20.00
Process is running for angle: 74.00
Process is running for angle: 56.00
Process is running for angle: 39.00
Process is running for angle: 3.00
Process is running for angle: 75.00
Process is running for angle: 21.00
Process is running for angle: 40.00
Process is running for angle: 57.00
Process is running for angle: 4.00
Process is running for angle: 76.00
Process is running for angle: 22.00
Process is running for angle: 41.00
Process is running for angle: 58.00
Process is running for angle: 23.00
Process is running for angle: 5.00
Process is running for angle: 77.00
Process is running for angle: 42.00
Process is running for angle: 24.00
Process is running for angle: 59.00
Process is running for angle: 6.00
Process is running for angle: 78.00
Process is running for angle: 43.00
Process is running for angle: 25.00
Process is running for angle: 60.00
Process is running for angle: 7.00
Process is running for angle: 79.00
Process is running for angle: 26.00
Process is running for angle: 8.00
Process is running for angle: 61.00
Process is running for angle: 44.00
Process is running for angle: 27.00
Process is running for angle: 45.00
Process is running for angle: 62.00
Process is running for angle: 9.00
Process is running for angle: 80.00
Process is running for angle: 28.00
Process is running for angle: 63.00
Process is running for angle: 46.00
Process is running for angle: 10.00
Process is running for angle: 81.00
Process is running for angle: 29.00
Process is running for angle: 64.00
Process is running for angle: 47.00
Process is running for angle: 11.00
Process is running for angle: 82.00
Process is running for angle: 30.00
Process is running for angle: 65.00
Process is running for angle: 48.00
Process is running for angle: 12.00
Process is running for angle: 31.00
Process is running for angle: 83.00
Process is running for angle: 66.00
Process is running for angle: 49.00
Process is running for angle: 13.00
Process is running for angle: 32.00
Process is running for angle: 67.00
Process is running for angle: 14.00
Process is running for angle: 84.00
Process is running for angle: 50.00
Process is running for angle: 33.00
Process is running for angle: 15.00
Process is running for angle: 68.00
Process is running for angle: 85.00
Process is running for angle: 51.00
Process is running for angle: 16.00
Process is running for angle: 34.00
Process is running for angle: 69.00
Process is running for angle: 86.00
Process is running for angle: 52.00
Process is running for angle: 35.00
Process is running for angle: 17.00
Process is running for angle: 70.00
Process is running for angle: 87.00
Process is running for angle: 90.00
Process is running for angle: 53.00
Process is running for angle: 108.00
Process is running for angle: 88.00
Process is running for angle: 71.00
Process is running for angle: 126.00
Process is running for angle: 109.00
Process is running for angle: 91.00
Process is running for angle: 89.00
Process is running for angle: 127.00
Process is running for angle: 144.00
Process is running for angle: 110.00
Process is running for angle: 162.00
Process is running for angle: 92.00
Process is running for angle: 128.00
Process is running for angle: 145.00
Process is running for angle: 111.00
Process is running for angle: 93.00
Process is running for angle: 163.00
Process is running for angle: 129.00
Process is running for angle: 112.00
Process is running for angle: 94.00
Process is running for angle: 164.00
Process is running for angle: 146.00
Process is running for angle: 130.00
Process is running for angle: 113.00
Process is running for angle: 147.00
Process is running for angle: 165.00
Process is running for angle: 95.00
Process is running for angle: 131.00
Process is running for angle: 148.00
Process is running for angle: 114.00
Process is running for angle: 96.00
Process is running for angle: 166.00
Process is running for angle: 132.00
Process is running for angle: 149.00
Process is running for angle: 115.00
Process is running for angle: 167.00
Process is running for angle: 97.00
Process is running for angle: 133.00
Process is running for angle: 150.00
Process is running for angle: 98.00
Process is running for angle: 116.00
Process is running for angle: 134.00
Process is running for angle: 168.00
Process is running for angle: 151.00
Process is running for angle: 135.00
Process is running for angle: 117.00
Process is running for angle: 99.00
Process is running for angle: 169.00
Process is running for angle: 152.00
Process is running for angle: 136.00
Process is running for angle: 100.00
Process is running for angle: 118.00
Process is running for angle: 170.00
Process is running for angle: 137.00
Process is running for angle: 153.00
Process is running for angle: 101.00
Process is running for angle: 119.00
Process is running for angle: 171.00
Process is running for angle: 154.00
Process is running for angle: 138.00
Process is running for angle: 102.00
Process is running for angle: 120.00
Process is running for angle: 155.00
Process is running for angle: 139.00
Process is running for angle: 172.00
Process is running for angle: 103.00
Process is running for angle: 121.00
Process is running for angle: 140.00
Process is running for angle: 156.00
Process is running for angle: 173.00
Process is running for angle: 104.00
Process is running for angle: 141.00
Process is running for angle: 122.00
Process is running for angle: 157.00
Process is running for angle: 174.00
Process is running for angle: 142.00
Process is running for angle: 158.00
Process is running for angle: 105.00
Process is running for angle: 123.00
Process is running for angle: 175.00
Process is running for angle: 159.00
Process is running for angle: 124.00
Process is running for angle: 106.00
Process is running for angle: 143.00
Process is running for angle: 125.00
Process is running for angle: 180.00
Process is running for angle: 176.00
Process is running for angle: 107.00
Process is running for angle: 160.00
Process is running for angle: 198.00
Process is running for angle: 181.00
Process is running for angle: 177.00
Process is running for angle: 161.00
Process is running for angle: 216.00
Process is running for angle: 178.00
Process is running for angle: 199.00
Process is running for angle: 182.00
Process is running for angle: 234.00
Process is running for angle: 217.00
Process is running for angle: 179.00
Process is running for angle: 183.00
Process is running for angle: 200.00
Process is running for angle: 235.00
Process is running for angle: 252.00
Process is running for angle: 218.00
Process is running for angle: 236.00
Process is running for angle: 184.00
Process is running for angle: 201.00
Process is running for angle: 253.00
Process is running for angle: 219.00
Process is running for angle: 185.00
Process is running for angle: 237.00
Process is running for angle: 254.00
Process is running for angle: 202.00
Process is running for angle: 186.00
Process is running for angle: 238.00
Process is running for angle: 220.00
Process is running for angle: 203.00
Process is running for angle: 255.00
Process is running for angle: 239.00
Process is running for angle: 187.00
Process is running for angle: 256.00
Process is running for angle: 240.00
Process is running for angle: 221.00
Process is running for angle: 188.00
Process is running for angle: 204.00
Process is running for angle: 257.00
Process is running for angle: 241.00
Process is running for angle: 189.00
Process is running for angle: 205.00
Process is running for angle: 222.00
Process is running for angle: 258.00
Process is running for angle: 190.00
Process is running for angle: 242.00
Process is running for angle: 223.00
Process is running for angle: 206.00
Process is running for angle: 259.00
Process is running for angle: 191.00
Process is running for angle: 224.00
Process is running for angle: 207.00
Process is running for angle: 243.00
Process is running for angle: 260.00
Process is running for angle: 225.00
Process is running for angle: 244.00
Process is running for angle: 208.00
Process is running for angle: 261.00
Process is running for angle: 192.00
Process is running for angle: 245.00
Process is running for angle: 209.00
Process is running for angle: 193.00
Process is running for angle: 226.00
Process is running for angle: 262.00
Process is running for angle: 246.00
Process is running for angle: 210.00
Process is running for angle: 194.00
Process is running for angle: 227.00
Process is running for angle: 263.00
Process is running for angle: 247.00
Process is running for angle: 211.00
Process is running for angle: 264.00
Process is running for angle: 195.00
Process is running for angle: 228.00
Process is running for angle: 248.00
Process is running for angle: 265.00
Process is running for angle: 212.00
Process is running for angle: 229.00
Process is running for angle: 196.00
Process is running for angle: 249.00
Process is running for angle: 266.00
Process is running for angle: 230.00
Process is running for angle: 197.00
Process is running for angle: 213.00
Process is running for angle: 250.00
Process is running for angle: 267.00
Process is running for angle: 231.00
Process is running for angle: 214.00
Process is running for angle: 251.00
Process is running for angle: 270.00
Process is running for angle: 268.00
Process is running for angle: 232.00
Process is running for angle: 288.00
Process is running for angle: 215.00
Process is running for angle: 271.00
Process is running for angle: 289.00
Process is running for angle: 233.00
Process is running for angle: 272.00
Process is running for angle: 269.00
Process is running for angle: 306.00
Process is running for angle: 290.00
Process is running for angle: 324.00
Process is running for angle: 273.00
Process is running for angle: 307.00
Process is running for angle: 342.00
Process is running for angle: 325.00
Process is running for angle: 291.00
Process is running for angle: 308.00
Process is running for angle: 274.00
Process is running for angle: 343.00
Process is running for angle: 326.00
Process is running for angle: 275.00
Process is running for angle: 309.00
Process is running for angle: 344.00
Process is running for angle: 292.00
Process is running for angle: 327.00
Process is running for angle: 310.00
Process is running for angle: 276.00
Process is running for angle: 345.00
Process is running for angle: 328.00
Process is running for angle: 293.00
Process is running for angle: 311.00
Process is running for angle: 277.00
Process is running for angle: 346.00
Process is running for angle: 294.00
Process is running for angle: 312.00
Process is running for angle: 329.00
Process is running for angle: 278.00
Process is running for angle: 295.00
Process is running for angle: 347.00
Process is running for angle: 313.00
Process is running for angle: 279.00
Process is running for angle: 330.00
Process is running for angle: 348.00
Process is running for angle: 280.00
Process is running for angle: 296.00
Process is running for angle: 331.00
Process is running for angle: 314.00
Process is running for angle: 349.00
Process is running for angle: 281.00
Process is running for angle: 332.00
Process is running for angle: 315.00
Process is running for angle: 297.00
Process is running for angle: 282.00
Process is running for angle: 333.00
Process is running for angle: 316.00
Process is running for angle: 350.00
Process is running for angle: 298.00
Process is running for angle: 334.00
Process is running for angle: 283.00
Process is running for angle: 317.00
Process is running for angle: 351.00
Process is running for angle: 335.00
Process is running for angle: 299.00
Process is running for angle: 284.00
Process is running for angle: 318.00
Process is running for angle: 336.00
Process is running for angle: 285.00
Process is running for angle: 352.00
Process is running for angle: 300.00
Process is running for angle: 337.00
Process is running for angle: 319.00
Process is running for angle: 286.00
Process is running for angle: 353.00
Process is running for angle: 301.00
Process is running for angle: 338.00
Process is running for angle: 320.00
Process is running for angle: 287.00
Process is running for angle: 354.00
Process is running for angle: 321.00
Process is running for angle: 339.00
Process is running for angle: 302.00
Process is running for angle: 355.00
Process is running for angle: 322.00
Process is running for angle: 340.00
Process is running for angle: 323.00
Process is running for angle: 303.00
Process is running for angle: 356.00
Process is running for angle: 341.00
Process is running for angle: 304.00
Process is running for angle: 357.00
Process is running for angle: 305.00
Process is running for angle: 358.00
Process is running for angle: 359.00
residuals (offsets): [ 0.01415679 0.02621993 -0.02744315 0.00635802 0.00837833 -0.0176962
-0.03312964 -0.0547925 -0.06124026 -0.07192343 -0.06382566 -0.07711383
-0.09335087 -0.06946001 -0.02784876 -0.01598752 0.02652482 0.0526809
0.10291377 0.07627605 0.08016999 0.12327355 0.04013548 -0.01765467
-0.05793429 -0.08913544 -0.07950251 -0.14093212 -0.1179294 -0.11975682
-0.10624286 -0.09006137 -0.08172908 -0.010413 -0.00552254 0.00274251
0.04476128 0.01594012 0.02296539 0.01412726 0.01801435 -0.01715912
-0.01237452 0.0053138 -0.01071325 0.03306639 0.03159408 0.07173099
0.05520899 0.05662432 0.06153248 0.04479955 0.0446696 0.05382092
0.04781707 0.042598 0.02748159 -0.0064816 -0.02805345 -0.0473227
-0.08893345 -0.08833836 -0.08534747 -0.07996837 -0.09014496 -0.03539202
-0.0025495 0.01278922 0.00407768 0.00029818 0.03154075 0.01252282
0.01683253 0.0227989 -0.00583583 0.01100947 0.01884958 0.01248943
0.01915697 -0.05120149 -0.02140614 -0.03594936 -0.01526049 -0.03755925
0.0020905 -0.01593293 0.02130371 0.070312 0.05976488 -0.10668152
-0.00836779 -0.11304176 -0.10291612 -0.0847637 -0.06208991 -0.04016928
-0.02742539 -0.01474581 -0.00775172 0.02092931 0.01610879 0.06267376
0.05770043 0.09734702 0.00707407 0.01588038 0.01558269 0.01845083
-0.00488014 -0.00121936 -0.06574346 -0.02716663 -0.06885915 -0.06067351
-0.06191396 -0.07934442 -0.04945264 -0.08910434 -0.08209316 -0.04183414
-0.01807385 0.01175321 0.00666549 -0.00122192 0.02550499 -0.00028513
0.04187227 0.08907192 0.05335619 0.04342524 0.03197163 0.03092187
0.03864068 0.0133347 0.03647415 0.00441657 0.00738352 0.01447402
-0.02056979 0.00775961 0.02699863 0.02653935 -0.00689883 -0.02857662
-0.0681384 -0.07254049 -0.07039994 -0.05247677 -0.05968222 -0.13525
-0.02422292 -0.00751036 0.0053182 0.005684 0.00820559 -0.01459568
-0.02557285 -0.04037477 -0.04252332 -0.00910463 0.01885289 0.03203826
0.02915053 0.03351121 0.01289627 0.00787485 -0.02108522 0.02623633
0.00720997 -0.0164661 -0.054297 -0.03577489 -0.03970628 -0.06265529
-0.08648492 -0.07015189 -0.02141407 -0.0391194 -0.01151127 0.02796464
0.07364988 0.03137258 0.05278229 0.0701776 0.04926231 0.03958866
0.00339508 0.00364629 -0.03372952 -0.09176793 -0.05655963 -0.01994561
-0.00081988 -0.01809907 0.01446584 0.0213347 0.02078129 0.05374183
0.07443977 0.04919889 0.05073581 0.03685402 0.05245267 0.01744649
0.01075707 -0.03964537 -0.00938504 -0.091538 -0.08889958 -0.08732591
-0.04722571 -0.08299886 -0.05376568 -0.08557094 -0.10031426 -0.02973594
-0.01399427 -0.0006416 0.01085913 0.04555631 0.06768609 0.10799103
0.08667012 0.09762593 0.08343457 0.09639241 0.03309279 0.04705858
0.01035518 0.04044133 -0.00523871 -0.00949436 -0.02331282 0.00687046
-0.02207876 0.00073402 -0.02137484 -0.02862403 -0.00435431 0.0183008
-0.01314301 -0.02711993 -0.02477116 -0.01770914 -0.02204705 0.04688026
-0.00128672 -0.02053155 -0.02137172 0.05978472 -0.00291652 -0.03014758
-0.043392 -0.05146947 -0.04366336 -0.04390729 -0.0538862 -0.08633528
-0.07257503 -0.08294154 -0.04085304 0.02510685 0.04047097 0.05553998
0.04117964 0.04762758 0.0329635 0.07814601 0.05136966 -0.00359316
0.00784139 0.00064708 -0.05084923 -0.02312229 -0.05319722 0.00466931
-0.02432471 -0.0307057 -0.03659312 -0.02017166 -0.02322855 -0.00077957
-0.03275365 -0.04335271 -0.05119144 -0.04840665 -0.02492606 0.00547774
0.02577521 0.01792894 0.0312105 0.03976592 -0.01507302 -0.02169466
-0.04680949 -0.05637433 -0.07007032 -0.06404906 -0.05324035 -0.03124574
0.07103666 0.06939683 0.10697873 0.12017969 0.1262887 0.07210945
0.03972485 0.01580822 -0.01087311 -0.04779305 -0.07189097 -0.04013429
-0.04546326 -0.07357915 -0.07971995 -0.08327568 -0.04381112 -0.03969979
-0.01690859 0.01851756 0.03218444 0.08892022 0.10278834 0.10282111
0.1128942 0.09037248 0.02641673 0.05790975 0.05155123 0.03062749
0.02377281 -0.0059859 -0.02860232 -0.0471336 -0.04118932 -0.04994412
-0.10183897 -0.08192172 -0.05706774 -0.04549874 -0.04630793 -0.0349439
0.00056419 0.02860427 0.04372081 0.03937523 0.08573526 0.10128295
0.13033904 0.12599876 0.08425184 0.04289406 0.01264203 -0.02488372
-0.01537808 -0.05600838 -0.08103007 -0.06362085 -0.04672134 -0.03742703] [ 2.05493388e-04 6.04974839e-02 7.16232778e-02 2.06756570e-02
3.45553329e-02 1.92471037e-02 -2.32913813e-02 1.55192938e-02
-2.93722726e-02 3.10598173e-02 8.63198766e-03 2.90645733e-02
-3.27498055e-02 -5.23303173e-03 -1.75859648e-02 -8.30606402e-03
-5.28911126e-02 1.65703643e-04 -3.80932469e-02 -1.77625031e-02
5.26298332e-03 -4.85192110e-02 -4.65957052e-03 7.35379817e-03
-2.67922802e-03 -3.04678616e-02 2.89508175e-02 5.69191216e-02
4.17120810e-02 2.46409731e-02 -4.07273975e-03 -2.65628304e-02
-4.31039354e-02 -1.02573956e-02 -4.25373032e-02 8.77247627e-04
-2.04247999e-02 2.24242928e-02 2.93163256e-02 2.31386136e-02
2.74719376e-02 8.74461596e-03 4.52435866e-02 5.28449234e-02
6.18166294e-02 6.29331827e-02 1.90016173e-02 5.61054945e-02
2.69496912e-02 -4.66243553e-02 -5.86095413e-02 -1.05482680e-01
-1.19592315e-01 -1.08115687e-01 -1.12022307e-01 -1.19345722e-02
6.55610091e-03 3.34147464e-02 9.83074596e-02 1.65846713e-01
1.19999316e-01 6.97314286e-02 9.52968109e-02 2.94811958e-02
1.90904939e-03 -1.42984619e-02 6.30100017e-03 -2.18761870e-02
-2.27671663e-02 -7.60810718e-02 -9.00612303e-02 -1.04476122e-01
-1.29084640e-01 -9.57215651e-02 -9.89813857e-02 -6.87266564e-02
-6.26495943e-02 -3.05948470e-02 8.88411359e-02 5.90883360e-02
9.75925570e-02 1.19157093e-01 1.76426755e-01 1.23322549e-01
1.39764944e-01 9.69149868e-02 7.45363466e-02 1.27751784e-02
-2.36503698e-02 3.78284983e-02 -6.60785721e-03 -6.71170210e-04
1.61306761e-02 1.55016014e-02 -3.03343699e-02 -2.41313595e-02
-2.84054245e-02 -5.61621427e-03 -9.25238023e-02 -1.07296119e-01
-6.19612697e-02 -9.65030815e-02 -1.01522292e-01 -4.78651420e-02
-5.70752039e-02 -5.65521283e-03 7.39053795e-03 6.83977352e-02
9.66212263e-02 1.27494073e-01 9.43545236e-02 8.78552156e-02
-2.23969709e-02 -2.59669164e-02 -4.30769612e-02 -1.24840265e-02
-5.94909209e-02 3.59357823e-03 6.78335018e-03 3.07936697e-03
2.35258207e-02 1.23430878e-02 5.76792673e-02 2.09977154e-03
4.80003309e-03 -2.71787796e-03 -2.31723418e-03 1.93435250e-03
-6.18435386e-02 -1.95651670e-02 2.06854004e-02 3.06625305e-02
6.85083196e-02 6.41265803e-02 7.20761643e-02 4.98761631e-02
8.11316417e-02 6.26437858e-02 5.70443169e-02 3.99353981e-02
-3.49108668e-03 1.51117591e-02 -5.31369987e-02 -1.19430537e-02
-2.72403310e-02 -7.20560612e-02 -9.11143555e-02 -7.65450942e-02
-7.14124478e-02 -9.52603426e-02 -1.29765872e-02 4.64988867e-02
4.09609937e-02 1.16925386e-01 8.30965761e-02 5.47176714e-02
7.14836073e-03 -5.93065020e-03 -5.01609196e-02 -5.00348031e-02
-4.09504998e-02 -6.96853766e-02 -7.33307064e-02 -2.37119879e-02
-3.60705962e-02 2.88672410e-02 6.55562275e-02 8.65962004e-02
7.61802725e-02 6.50833167e-02 7.21513156e-02 1.64936488e-02
-2.33624668e-02 -1.00912276e-01 -9.56862695e-02 -1.87053949e-01
-1.33937856e-01 -1.25419728e-01 -6.08878726e-02 -9.12578580e-04
9.58173591e-02 9.72355132e-02 4.45408542e-02 8.24429399e-02
2.26852582e-02 -4.85196290e-02 -5.31300575e-02 -9.17620659e-02
-6.26828416e-02 -1.92985451e-02 2.27242820e-03 -1.56085346e-03
1.58602765e-03 2.30553261e-02 -2.08149243e-02 -2.17367849e-02
1.39671825e-02 4.07008678e-02 3.20427448e-02 4.96780100e-02
5.64531815e-02 2.56404747e-02 -8.51756310e-03 -3.54465287e-02
-2.15608865e-02 -3.96242030e-02 -4.43117180e-02 -2.94692622e-02
2.66772245e-02 8.68218105e-03 1.09366250e-02 4.40007191e-02
1.44068313e-02 -2.36362561e-03 1.12481041e-02 -2.22105210e-02
-5.28326512e-04 -1.67062943e-03 2.03506550e-02 3.09540000e-02
6.10490011e-03 -1.01708684e-02 1.01242533e-02 -4.13254166e-02
3.90430900e-02 -1.70258384e-02 4.71413479e-02 1.08167448e-01
4.61350541e-02 2.96258289e-02 1.84822293e-03 -9.97787449e-03
-5.75858778e-02 -6.45301042e-02 -6.14452999e-02 7.65049648e-03
-4.71862879e-03 1.05377753e-02 3.63709532e-02 6.16213339e-02
-2.91688245e-02 -5.74455585e-03 -8.33193831e-02 -3.75099652e-02
-4.72396969e-02 -1.04130178e-01 -4.05438334e-02 2.00135434e-02
1.22824087e-02 5.51516073e-02 5.33222329e-02 4.77979757e-02
3.06245766e-02 -2.61491038e-02 -4.25993003e-02 -8.93460193e-02
-1.17678504e-01 -9.34718876e-02 -5.48994676e-02 -6.77085727e-03
5.66908219e-02 6.84447654e-02 1.51849104e-01 1.65723382e-01
1.66747022e-01 1.30894321e-01 4.67500284e-02 -3.85316731e-02
-8.68034087e-02 -1.96720875e-01 -2.38741605e-01 -2.97476140e-01
-2.91610561e-01 -2.01892792e-01 -1.28239104e-01 -3.28329593e-02
-3.84044494e-04 5.63604887e-02 1.35954880e-01 9.76392244e-02
3.83744105e-02 -3.64882659e-02 -9.73311400e-03 2.18049653e-03
1.74708195e-01 1.81559025e-01 1.68832002e-01 1.09019844e-01
1.62978674e-01 8.11516010e-02 -8.80464523e-03 -7.25744212e-02
-1.03763354e-01 -1.30764063e-01 -1.87086899e-01 -1.33382870e-01
-1.49462489e-01 -9.20128829e-02 -1.37168284e-02 7.55218518e-02
1.29430112e-01 9.26455959e-02 1.22828482e-01 9.60202658e-02
7.43460292e-02 3.92982700e-02 1.57284496e-02 -3.69944074e-02
-2.85063330e-02 -4.60111090e-02 -5.65891514e-02 -7.28057920e-02
-5.86502717e-03 -1.02146729e-02 -2.52298797e-02 -1.67538191e-02
3.09056103e-02 5.25300091e-02 5.23660981e-02 3.03785577e-02
1.97978015e-02 3.33030492e-02 -3.97623237e-02 -2.83950493e-02
-5.73528341e-02 -3.95601424e-02 -4.93320029e-03 4.25664342e-02
6.91811859e-02 6.24308040e-02 5.47281020e-02 2.52655891e-02
1.01527918e-01 5.71339791e-02 1.37288919e-02 5.51915656e-03
-2.81605563e-02 -3.63519556e-02 -2.68342149e-02 -7.15562470e-02
-6.91640744e-02 -6.37262523e-02 -4.48211447e-02 -3.59883383e-02
-3.32283364e-03 1.69119399e-02 1.01642725e-01 7.63727306e-02
5.10312472e-02 6.66624678e-02 4.81481261e-02 -2.14890225e-02
-4.61750684e-02 -1.53899236e-01 -1.62938416e-01 -1.60059537e-01
-1.24157290e-01 -5.16888071e-02 3.55256468e-02 1.25062180e-01] [-1.21094502e+01 -8.53314459e+00 -5.09504996e+00 -4.16540942e+00
3.80800282e+00 -4.01162122e+00 -2.58607612e+00 2.90949736e+00
-3.65808317e+00 1.49709375e+00 7.25925310e-01 6.44362045e+00
8.53047555e+00 2.68775735e+00 -1.63438283e+00 2.56596511e-01
-3.27105765e+00 -3.80827622e+00 -8.77792128e+00 -5.89188351e-01
-2.08514212e+01 -1.26333483e+01 -7.76610835e+00 -8.16203630e+00
-9.41955265e+00 -3.28045987e+00 -5.88614463e+00 3.20820966e+00
4.31933513e+00 3.51312656e+00 3.57244363e+00 4.36994112e+00
6.55267248e-02 -2.78402831e+00 -5.30928853e+00 -4.40840418e+00
-4.64106838e+00 -7.42368561e-01 9.44022716e-01 4.60236224e+00
-2.65330405e+00 2.66404348e+00 1.35565785e+00 2.54978188e+00
1.14594284e+01 7.70262624e+00 1.14311991e+01 5.79972913e+00
9.83069302e+00 9.55237429e+00 6.51398763e+00 -3.14110119e+00
-5.91166787e+00 -1.05823710e+01 -1.89473597e+01 -1.92793801e+01
-1.94548765e+01 -2.23727700e+01 -1.47161534e+01 -8.39146146e+00
-6.42503124e+00 -5.55588719e+00 1.18938117e+00 3.15928541e+00
8.91336681e+00 1.20364406e+01 1.36588495e+01 7.35625462e+00
6.65346998e+00 7.93599811e+00 1.37639204e+00 -3.06885637e+00
-3.78357987e+00 -9.92363988e+00 -1.25296781e+01 -1.79770753e+01
-2.23295513e+01 -2.27602269e+01 -1.94338360e+01 -2.48995357e+01
-1.84168887e+01 -1.46425445e+01 -1.16227359e+01 2.99214548e+00
5.90545511e+00 1.09025565e+01 1.82924616e+01 2.32877948e+01
2.07728266e+01 2.96142266e+01 1.77156586e+01 2.29799810e+01
1.86306429e+01 1.70869328e+01 1.74887153e+01 9.96682782e+00
9.52666501e+00 6.31642626e+00 7.50940350e+00 -4.26207497e-01
-4.58937419e-01 -5.54503542e+00 -1.84967566e+01 -2.12977257e+01
-6.86098724e+00 -1.58836691e+01 -1.14736862e+01 -1.04951343e+01
-1.18119664e+01 -3.34908646e+00 -1.74536915e+00 2.56097411e+00
5.65972477e+00 1.30720362e+00 -1.09741870e+00 -4.76757115e+00
-6.17211008e+00 -5.46497397e+00 -8.04485430e+00 -5.39720549e+00
-6.56030975e+00 -3.34851673e+00 -3.75668547e+00 8.74473361e-01
-1.75542294e-01 6.98547628e-02 -1.14287924e-01 -4.07192882e+00
2.07959197e-01 -3.66031946e+00 -6.92319155e+00 -7.35109813e+00
-3.61466822e+00 -4.08756093e-01 2.03361058e+00 -7.26198756e-03
7.17499353e+00 1.37646925e+01 1.18330210e+01 1.30276836e+01
1.61500373e+01 1.40846097e+01 1.67529247e+01 7.47332408e+00
7.05384106e+00 7.94431805e+00 2.61219643e+00 5.24974515e-01
-4.53333140e+00 -7.41270641e+00 -6.13385365e+00 -5.65478148e+00
3.09263606e-01 8.55919100e+00 9.52046439e+00 9.34649227e+00
1.54582137e+01 9.69359984e+00 1.21390077e+01 9.07146043e+00
7.44436035e+00 3.05533087e+00 2.38569733e+00 1.13457939e+00
-1.72619516e+00 2.88540164e+00 4.18187596e+00 4.09147382e+00
8.21734667e+00 1.11445564e+01 1.45000086e+01 1.53796039e+01
1.93117418e+01 1.58147065e+01 7.03483866e+00 9.47412817e-01
-1.19864976e+00 -5.04100377e+00 -9.09048854e+00 -7.61240627e+00
-6.24904961e-01 -8.42576927e-01 4.82036454e+00 2.92800624e+00
8.05300591e+00 6.69481128e+00 3.88755010e+00 2.97170093e+00
-9.11957759e-01 1.20251576e+00 -3.77378524e+00 -1.20789054e+00
-1.71407543e+00 1.99134214e+00 -4.66361313e+00 -3.56302510e+00
-4.86437130e+00 -3.53988444e+00 -2.80273628e+00 -1.42455142e-01
2.56562804e+00 3.24960734e+00 1.76697696e+00 4.51116296e+00
1.57157718e+00 6.76014360e-01 -2.43324937e+00 1.89952226e-01
-4.59418855e-01 -4.55146840e-01 6.02340770e-01 -3.89669393e+00
-4.13642129e+00 -4.70588622e+00 -1.05189714e+00 -7.08195662e+00
-6.03905799e+00 -1.77497831e+00 -7.74388409e+00 -7.13797650e+00
-3.59167495e+00 2.14427607e+00 1.29563719e+00 3.90518090e+00
-1.55031426e+00 -5.41152641e+00 7.33775907e-01 5.08183048e+00
6.89602262e+00 1.15567733e+01 9.73465896e+00 1.14992231e+01
3.94873921e+00 3.70109196e+00 1.61261889e-01 3.55515899e+00
1.90833923e-01 -4.28899480e+00 3.77238013e+00 1.38779003e-01
1.50518153e+00 1.38284020e+00 -3.99633392e+00 -6.38310715e-01
-4.46476139e+00 -7.58551455e+00 -1.10469907e+01 -1.03960433e+01
-1.20642985e+01 -5.93114641e+00 -7.14002492e+00 -3.65567257e+00
1.07528561e+00 3.19280876e+00 1.88799311e-01 -1.76619358e+00
-2.58381870e+00 -5.70951818e+00 -1.26911106e+01 -1.74816865e+01
-1.54691356e+01 -4.17283749e+00 -4.56716813e+00 3.79517682e+00
1.47033388e+01 1.88177393e+01 3.03573809e+01 3.27176710e+01
2.87719033e+01 1.58790273e+01 1.42920667e+01 2.88426153e+00
-5.38904828e+00 -1.95341328e+01 -2.69500694e+01 -3.58998028e+01
-3.56150098e+01 -3.48770697e+01 -2.95209025e+01 -2.44418149e+01
-1.90106618e+01 -1.29884303e+01 -1.32896876e+01 -1.37633355e+01
-1.58640191e+00 1.14265847e+00 8.32991434e-01 7.65837285e+00
1.98298397e+01 2.04443344e+01 2.42223121e+01 2.64397793e+01
1.61253360e+01 5.99931660e+00 8.05504478e+00 -1.46068933e+00
-1.05120015e+01 -1.81141936e+01 -1.78638026e+01 -2.21460416e+01
-1.12831033e+01 -3.03359957e+00 -8.60170804e+00 3.53843663e+00
9.45959483e+00 1.23540707e+01 1.60205826e+01 1.21770432e+01
1.39099792e+01 9.12822226e+00 4.23744792e+00 3.40275508e+00
7.40198195e+00 -8.14393648e+00 -1.97205326e+00 -2.10991289e+00
-1.98972326e+00 -3.53068061e-01 3.89718037e+00 5.42286422e+00
4.46807360e+00 6.65133986e+00 4.21690880e+00 -2.34376433e-01
-1.52556079e+00 -5.66808346e+00 -9.38940910e+00 -8.05851200e+00
-2.10745352e+00 1.94853571e+00 4.82151837e+00 3.81192847e+00
5.52092651e+00 6.77197003e+00 1.02450717e+01 1.10120179e+01
1.05944210e+01 9.09138013e+00 6.31333637e+00 7.31619540e+00
3.66622994e+00 5.47017671e-01 -2.88522384e+00 1.94019551e+00
-2.13308100e+00 -4.31186534e+00 -2.00187660e-01 5.25415444e+00
8.41102525e+00 9.18822781e+00 9.93595353e+00 9.65497491e+00
1.21510897e+01 3.29332988e+00 -1.81935773e+00 -1.05679625e+01
-1.92903963e+01 -2.28660510e+01 -1.96646716e+01 -1.58680286e+01]
[[ 1.41567882e-02 2.05493388e-04 -1.21094502e+01]
[ 2.62199280e-02 6.04974839e-02 -8.53314459e+00]
[-2.74431538e-02 7.16232778e-02 -5.09504996e+00]
...
[-6.36208519e-02 -5.16888071e-02 -2.28660510e+01]
[-4.67213445e-02 3.55256468e-02 -1.96646716e+01]
[-3.74270297e-02 1.25062180e-01 -1.58680286e+01]]
(360, 3)
percentage for r: 68.6111111111111%
percentage for theta: 69.72222222222221%
percentage for f: 70.83333333333334%
Confidence intervals:
r: -0.024134815678862734 [-0.07158544342526864,0.07912075325950745]
theta: 0.00192083834674861 [-0.07318592808734081,0.07318592808734081]
f: -0.6380453955114689 [-10.4832251622406,10.4832251622406]
Gaussian fit results:
r: -0.005339318215964717 +-0.05274467671476413
theta: -0.0007472801152008336 +-0.07271145216585939
f: -0.001480924817492103 +-10.612436375468306
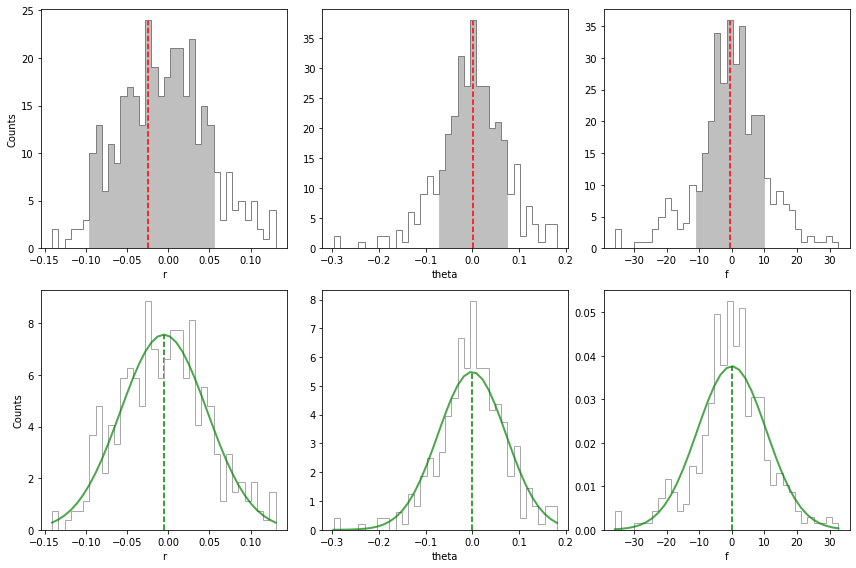
Again, if you wish to write the result (to avoid having to run the previous box again), just set write=True. This will pickle the result:
[45]:
write=False
if write:
output = {'speckle_res':speckle_res}
with open('../datasets/my_speckle_residuals_results', 'wb') as fileSave:
pickle.dump(output, fileSave)
Load pickled results from disk:
[46]:
with open('../datasets/speckle_residuals_results','rb') as fi:
myPickler = pickle.Unpickler(fi)
sp_unc_result = myPickler.load()
print(sp_unc_result.keys())
speckle_res = sp_unc_result['speckle_res']
sp_unc, mean_dev, p_simplex, offset, chi2, nit, success = speckle_res
dict_keys(['speckle_res'])
The speckle uncertainty associated to each parameter is contained in sp_unc which corresponds to the 1\(\sigma\) width of a Gaussian distribution fitted to the offset distribution (i.e. the differences with respect to injected ground truths):
[47]:
sp_unc
[47]:
array([ 0.05350939, 0.07376633, 10.64773837])
For comparison, the uncertainties found by the MCMC procedure were:
[48]:
sigma
[48]:
array([ 0.13676103, 0.15744912, 21.66963444])
5.3.5. Final uncertainties¶
The final uncertainties on the planet parameters should include both statistical and systematic uncertainties. The former include both the photon noise and residual speckle noise uncertainties discussed above. The latter include both the uncertainty on the star location (which may be non-negligible when using a coronagraph) and instrumental calibration errors, including:
- the uncertainty on the plate scale (for \(r\), when converting to angular separation) - note that it is proportional to the radial separation itself;
- the uncertainty on the PA of true north (for \(\theta\)).
The uncertainty on the star location is of the order of 0.3px in individual NACO+AGPM images (this is the typical precision by manual recentering during the observation). Given the shift plots in Tutorial 2, it appears the autocorrelation timescale is of the order of ~5 frames. Therefore, considering that there are 61 frames in the datacube, the uncertainty on the star location in the final combined image must be roughly:
[49]:
cen_unc_indiv = 0.3 #px
cen_unc = cen_unc_indiv/np.sqrt(61/5) #px
cen_unc
[49]:
0.08588975014708022
The latter can be translated into an uncertainty on \(\theta\) by division by the radial separation of the companion. The stellar centering uncertainties on each planet parameter can thus be expressed as:
[50]:
star_unc = np.array([cen_unc*pxscale_naco,
np.rad2deg(cen_unc/val_max['r']),
0]) # uncertainty on each of the 3 parameters due to stellar centering
where the multiplication by pxscale_naco has converted the radial separation in arcsec.
For the instrumental calibration errors, we adopt the values quoted in Absil et al. (2013). Note that the uncertainty related to the plate scale is directly proportional to the radial separation of the companion.
[51]:
dr_unc = 0.00004 # plate scale uncertainty in arcsec per px
tn_unc = 0.09 # deg
syst_unc = np.array([val_max['r']*dr_unc,
tn_unc,
0])
The final uncertainties are then the different sources of uncertainty added quadratically - after conversion of the radial separation to arcsec:
[52]:
sigma[0] *= pxscale_naco
sp_unc[0] *= pxscale_naco
if mu_sigma:
final_unc = np.sqrt(np.power(sigma,2) + np.power(star_unc,2) + np.power(syst_unc,2))
else:
final_unc = np.sqrt(np.power(sigma,2) + np.power(sp_unc,2) + np.power(star_unc,2) + np.power(syst_unc,2))
[53]:
msg = "The final estimates on the radial separation, PA and flux of the companion, including uncertainties, are: \n"
msg+= "r = {:.2f}+-{:.2f} mas (GT: {:.2f} mas), \n"
msg+= "PA = {:.2f}+-{:.2f} deg (GT: {:.2f} deg) \n"
msg+= "f = {:.2f}+-{:.2f} ADUs (GT: {:.2f} ADUs)"
print(msg.format(val_max['r']*pxscale_naco*1000, final_unc[0]*1000, rad_fc*pxscale_naco*1000,
val_max['theta'], final_unc[1], theta_fc,
val_max['f'], final_unc[2], flux_fc))
The final estimates on the radial separation, PA and flux of the companion, including uncertainties, are:
r = 831.33+-4.56 mas (GT: 829.29 mas),
PA = 239.92+-0.24 deg (GT: 240.00 deg)
f = 390.77+-21.67 ADUs (GT: 400.00 ADUs)
Let’s consider the Gaussian fit instead:
[54]:
msg = "Considering a Gaussian fit to the posterior distributions, the final estimates on the radial separation, PA and flux of the companion, including uncertainties, are: \n"
msg+= "r = {:.2f}+-{:.2f} mas (GT: {:.2f} mas), \n"
msg+= "PA = {:.2f}+-{:.2f} deg (GT: {:.2f} deg) \n"
msg+= "f = {:.2f}+-{:.2f} ADUs (GT: {:.2f} ADUs)"
print(msg.format(mu[0]*pxscale_naco*1000, final_unc[0]*1000, rad_fc*pxscale_naco*1000,
mu[1], final_unc[1], theta_fc,
mu[2], final_unc[2], flux_fc))
Considering a Gaussian fit to the posterior distributions, the final estimates on the radial separation, PA and flux of the companion, including uncertainties, are:
r = 830.00+-4.56 mas (GT: 829.29 mas),
PA = 239.95+-0.24 deg (GT: 240.00 deg)
f = 394.13+-21.67 ADUs (GT: 400.00 ADUs)